ANTISENSE RNA
(August 2003)
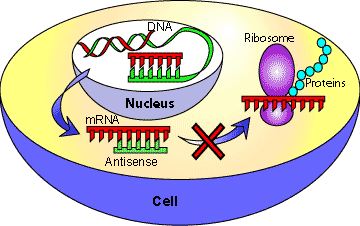
Messenger RNA (mRNA) is a single stranded molecule that is used as the template for protein translation. It is possible for RNA to form duplexes, similar to DNA, with a second sequence of RNA complementary to the first strand. This second sequence is called antisense RNA (Figure 1). The formation of double stranded RNA can inhibit gene expression in many different organisms including plants, flies, worms and fungi.
Co-Suppression
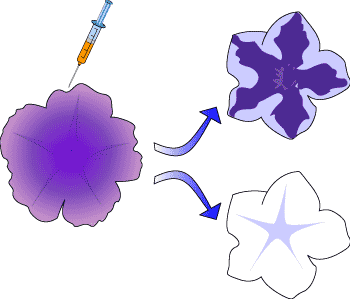
The first discovery of this inhibition in plants was more than a decade ago and occurred in petunias. Researchers were trying to deepen the purple colour of the flowers by injecting the gene responsible into the petunias but were surprised at the result. Instead of a darker flower, the petunias were either variegated (Figure 2) or completely white!
This phenomenon was termed co-suppression, since both the expression of the existing gene (the initial purple colour), and the introduced gene (to deepen the purple) were suppressed. Co-suppression has since been found in many other plant species and also in fungi. It is now known that double stranded RNA is responsible for this effect.
aRNA and RNAi
When antisense RNA (aRNA) is introduced into a cell, it binds to the already present sense RNA to inhibit gene expression. So what would happen if sense RNA is prepared and introduced into the cell? Since two strands of sense RNA do not bind to each other, it is logical to think that nothing would happen with additional sense RNA, but in fact, the opposite happens! The new sense RNA suppresses gene expression, similar to aRNA. While this may seem like a contradiction, it can be easily resolved by further examination. The cause is rooted in the prepared sense RNA. It turns out that preparations of sense RNA actually contain contaminating strands of antisense RNA. The sense and antisense strands bind to each other, forming a helix. This double helix is the actual suppressor of its corresponding gene. The suppression of a gene by its corresponding double stranded RNA is called RNA interference (RNAi), or post-transcriptional gene silencing (PTGS). The gene suppression by aRNA is likely also due to the formation of an RNA double helix, in this case formed by the sense RNA of the cell and the introduced antisense RNA.
How Does it Work?
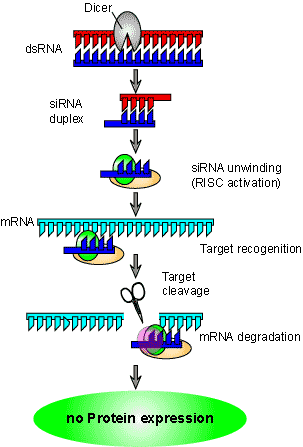
But how does the double stranded RNA cause gene suppression? Since the only RNA found in a cell should be single stranded, the presence of double stranded RNA signals is an abnormality. The cell has a specific enzyme (in Drosophila it is called Dicer) that recognizes the double stranded RNA and chops it up into small fragments between 21-25 base pairs in length. These short RNA fragments (called small interfering RNA, or siRNA) bind to the RNA-induced silencing complex (RISC). The RISC is activated when the siRNA unwinds and the activated complex binds to the corresponding mRNA using the antisense RNA. The RISC contains an enzyme to cleave the bound mRNA (called Slicer in Drosophila) and therefore cause gene suppression. Once the mRNA has been cleaved, it can no longer be translated into functional protein (Figure 3 and see a Flash animation of PTGS here).
Uses
The suppression of protein synthesis by introducing antisense RNA into a cell is very useful. A gene encoding the antisense RNA can be introduced fairly easily into organisms by using a plasmid vector or using a gene gun that shoots microscopic tungsten pellets coated with the gene into the plant cells. Once the antisense RNA is introduced, it will specifically inhibit the synthesis of the target protein by binding to mRNA. This is a quick way to create a knockout organism to study gene function. Using antisense RNA as a tool in this way is an exciting prospect for many molecular biologists.
Antisense RNA is also being investigated for use in cancer therapy. Injecting aRNA that is complementary to the proto-oncogene BCL-2 may be useful for treating some B-cell lymphomas and leukemias. Antisense oligodeoxynucleotides (ODNs) are also being studied for human therapy. ODNs are similar to antisense RNA, but they are made synthetically and are deoxynucleotides (like those in DNA) rather than nucleotides. ODNs are being tested for their effectiveness against HIV-1, cytomegalovirus (a member of the herpesvirus group), asthma and certain cancers.
Antisense RNA methods have also been used for commercial food production. You may have heard of the Flavr Savr tomato. This tomato was developed by Calgene Inc. of Davis, California in 1991 and was approved by the U.S. FDA in 1994. The tomato was the first whole food created by biotechnology that was evaluated by the FDA. One of the problems associated with tomato farming is that the fruit must be picked while still green in order to be shipped to market without being crushed. The enzyme that causes softening in tomatoes is polygalacturonase (PG). This enzyme breaks down pectin as the tomato ripens, leading to a softer fruit. Calgene suppressed the expression of the gene encoding PG by introducing a gene encoding the antisense strand of the mRNA. When the introduced gene was expressed, the antisense strand bound to the PG mRNA, suppressing the translation of the enzyme. The Flavr Savr tomatoes therefore had low PG levels and remained firmer when ripe. This meant the Flavr Savr tomatoes can ripen on the vine and then be shipped to market. Although the Flavr Savr tomatoes were approved for sale in the U.S., production problems and consumer wariness stopped the production of this fruit in 1997.
RNA interference is a field that was stumbled upon by accident while trying to improve the colour of petunias, however its implications may be far reaching in the near future.
Additional Readings
1. Kimball’s Biology Pages — Antisense RNA
2. Ambion — The RNA interference resource.
(Art by Fan Sozzi)