ATTACK OF THE SUPERBUGS: ANTIBIOTIC RESISTANCE
In the past 50 years, antibiotics have been critical in the fight against many diseases and infections. Their discovery was one of the leading causes for the dramatic rise of average life expectancy in the 20th century and their significance to public health would be impossible to overstate. Antibiotics are defined as any compound which either kills or severely impedes the growth of bacteria. Upon the introduction of penicillin into general clinical practice in 1944, formerly deadly illnesses such as Strep throat and tuberculosis became instantly curable. Today, our dependence on antibiotics is absolute. In 1998, in the United States, it was estimated that there were 80 million prescriptions of antibiotics for human use, the equivalent of about 12,500 tons in one year. When animal and agricultural uses of antibiotics are added to human use, it is estimated that in the past 50 years, more than 1 million tons have been produced and disseminated.
Almost as soon as antibiotics were introduced into clinical circulation, cases where their ability to effectively stop infection were observed. As the use of antibiotics became more widespread, the prevalence of antibiotic resistant bacteria increased. In a recent study in Atlanta, 25% of bacterial pneumonia cases were shown to be resistant to penicillin, while a further 25% of cases were resistant to more than one antibiotic. Resistance development has resulted in perpetual research and development in the search of new antibiotics in order to maintain a pool of effective drugs at all times. While the development of resistant strains is inevitable, the speed and scale of development has been exacerbated by the practices through which we use and disseminate antibiotics.
The discovery and development of antibiotics
In 1928, Alexander Fleming was searching for potential antibacterial compounds. He noticed that a patch of the mould Penicillium notatum had grown on a plate containing the bacteria Staphylococcus and that around the mould there was a zone where no Staphylococcus could grow. After more research, he was able to show that culture broth of the mold prevented growth of the Staphylococcus even when diluted up to 800 times. He named the active substance penicillin but was unable to stably isolate it. Several years later, in 1939, Ernst Chain and Howard Florey developed a way to isolate penicillin and used it to treat bacterial infections during the Second World War. The new drug came into clinical circulation in 1944 and made a huge impact on public health. For these discoveries Fleming, Chain and Florey were awarded the Nobel Prize for Medicine in 1945. Their discovery and development revolutionized modern medicine and paved the way for the development of many more natural antibiotics.
While Fleming was working on penicillin, Gerhard Domagk, a German doctor, announced the discovery of a synthetic molecule with antibacterial properties. The drug was named Prontosil and was the first of a long series of synthetic antibiotics called sulfonamides or sulfa drugs. Prontosil was introduced to clinical use in the 1930s and was used to combat urinary tract infections, pneumonia and other conditions. While sulfa drugs in many cases are not as effective as natural antibiotics, they are now in widespread use for the treatment of many conditions. Gerhard Domagk was awarded the Noble prize in 1939 for his discovery of prontosil.
Several strategies are currently used to find new antibacterial compounds. Potential candidates for natural antibiotics are discovered by screening bacterial and fungal species for antimicrobial activity. Semi-synthetics are made by derivitizing natural antibiotics. Synthetic drugs are serendipitously discovered or designed intelligently to bind to bacteria in a specific way.
Antibiotics generally work in one of five ways. These are:
- Inhibition of nucleic acid synthesis (e.g. Rifampicin; Chloroquine)
- Inhibition of protein synthesis (e.g. Tetracyclines; Chloramphenicol)
- Action on cell membrane (e.g. Polyenes; Polymyxin)
- Interference with enzyme system (e.g. Sulphamethoxazole)
- Action on cell wall (e.g. Penicillin; Vancomycin)
For example, penicillin works by blocking the formation of peptide bonds in the bacterial cell wall and thereby weakens it, leaving the bacterium susceptible to osmotic lysis. Although there appears to be wide assortment of antibiotics and mechanisms of action, there are a very limited number of targets through which bacteria are susceptible to antibacterial activity. Researchers must constantly look for new ways to combat bacteria.
Development and acquisition of resistance
Many infectious diseases have been brought under control around the world yet this remains the leading cause of death in the world. Furthermore, previously controlled infections are becoming increasingly common in patients with diseases like AIDS where the immune system is compromised. The microbes responsible for these infections are often antibiotic resistant pathogens. The ability for the pathogens to grow despite the presence of antibiotics, through the development of antibiotic resistance, has rendered victims as vulnerable as patients from the pre-antibiotic era. The development of resistance is inevitable following the introduction of a new antibiotic. In fact initial rates of resistance to new drugs are normally on the order of 1%. However modern uses of antibiotics have caused a huge increase in the number of resistance bacteria. In fact within eight to twelve years, after wide spread use, strains resistant to multiple drugs become widespread.
How do bacteria become resistant to antibiotics and what are the biochemical mechanisms that they use? Several mechanisms have been developed by bacteria in order to deal with antibiotics but all require either the modification of existing genetic material or the acquisition of new genetic material.
Originally it was believed that all resistance was acquired through spontaneous mutation. Development of resistance through this method is called primary resistance. Errors in DNA synthesis during replication and occasional failures in the DNA repair systems result in a spontaneous mutation frequency for an individual base pair of about 10-7-10-8. This means that for every 107-108 bacteria, we would expect one single base pair to be changed. Mutation is a very rare event. However, the spontaneous mutation rate to acquire a mutation that causes resistance is often even lower since multiple mutations must take place before primary antibiotic resistance can be acquired. In E. coli, it has been estimated that primary streptomycin resistance is acquired at a rate of approximately 10-9 when exposed to high concentrations of streptomycin. While this is an extremely rare event, the very fast growth rate of bacteria means that it doesn’t take long before resistance is developed in a population. Once the resistance genes are acquired, the genes can be transferred directly to all the bacteria’s progeny. This is known as vertical gene transfer.
The widespread development of multi-drug resistance in many species of bacteria simultaneously led scientists to believe that another mechanism beyond spontaneous mutation was responsible for the acquisition of antibiotic resistance. Lateral or horizontal gene transfer (HGT) is a process whereby genetic material contained in small packets of DNA can be transferred between individual bacteria. There are three possible mechanisms of HGT. These are transduction, transformation or conjugation. Transduction occurs when bacteria-specific viruses or bacteriophages transfer DNA between two closely related bacteria. Transformation is a process where parts of DNA are taken up by the bacteria from the external environment. This DNA is normally present in the external environment due to the death of another bacterium. Conjugation occurs when there is direct cell-cell contact between two bacteria (which need not be closely related) and transfer of small pieces of DNA called plasmids takes place. This is thought to be the main mechanism of antibiotic resistant gene transfer. Figure 1 shows the three mechanisms of HGT.
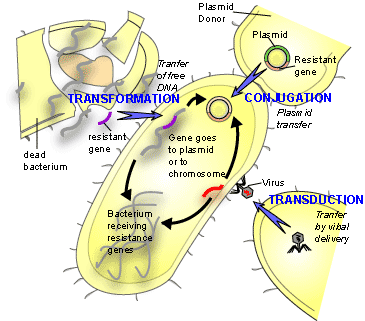
Mechanisms of antibiotic resistance
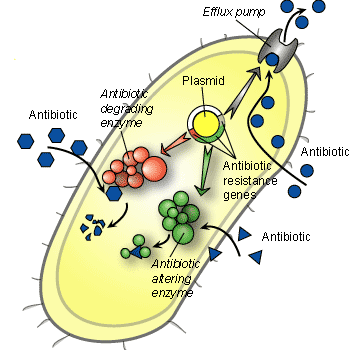
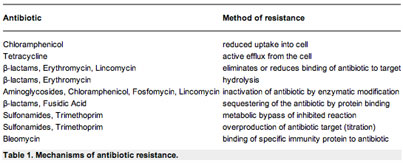
Several mechanisms have evolved in bacteria which confer them with antibiotic resistance. These mechanisms can either chemically modify the antibiotic or alternatively render it inactive through physical removal from the cell, or through modification of its target site. Figure 2 shows several different mechanisms of antibiotic resistance. The most common mode is enzymatic inactivation of the antibiotic. An existing enzyme is modified to process the antibiotic and modifies it so that it no longer affects the microorganism. An alternative strategy utilized by bacteria is the alteration of the antibiotic target site. Table 1 shows examples of several different methods of antibiotic resistance.
The scope of the problem
To combat the occurrence of resistant bacteria, pharmaceutical companies must constantly research, develop and test new antimicrobials in order to maintain a pool of effective drugs on the market. Five years ago, there were approximately 150 antibiotics available to the public with new drugs appearing every 8-10 years. This appears to be a substantial amount. However, these numbers are misleading as many of these drug targets are similar. Since the drug development process is very costly, pharmaceutical companies often concentrate on finding antimicrobials similar to the ones already found to reduce the risk of producing an unmarketable drug. This means that it is easy for microorganism to develop resistance to a similar drug to which it already has resistance. Past and current strategies to combat resistance are not effective.
The rate of resistance acquisition is accelerated by the widespread misuse of antibiotics. Antibiotics in animal feed, unnecessary and unfinished antibiotic prescriptions have been identified as causes for an enhanced rate of resistance development. Delivering antibiotics to livestock in animal feed is similar to giving people antibiotics all their life even when they are healthy. Food borne pathogens such as Salmonella, E. coli and Campylobacter are all bacteria that live in a symbiotic relationship with cows and chickens. They can acquire resistant genes, infect humans, cause food poisoning from consumption of beef or chicken and can horizontally transfer those resistance genes to other bacteria. Unnecessary prescriptions of antibiotics are made when antibiotics are prescribed for viral infections (antibiotics have no effect on viruses). This gives the opportunity for benign bacteria to acquire resistance that can be later passed on to pathogens. Unfinished antibiotic prescriptions leave some bacteria alive and exposes them to sub-inhibitory concentrations of antibiotics for a prolonged period of time. Mycobacterium tuberculosis is a slow growing bacteria which infects the lung and causes tuberculosis. This disease kills more adults than any other infectious disease. Yet at one time, health care workers believed they had the disease under control. Unfortunately due to the slow growing nature of the infection, treatment programs last for months or even years. This has led to many cases on unfinished prescriptions and 5% of strains now observed are completely resistant to all known treatments and hence incurable.
Combating antibiotic resistance
Several possible solutions have been proposed to not only slow down resistance but to also to reverse it. A decrease in the number of prescription for antibiotics in small children is occurring. The rate has been reduced by 23 % from 1995-2000 in children less than 3 yrs old with similar trends being mimicked in older children. Large scale public health education efforts are now underway to stress the importance of finishing prescriptions. Indeed in many places, failure to finish tuberculosis prescriptions can result in jail time. In addition, several countries such as the UK have regulations concerning the use of antibiotics in animal feed.
Reversal of resistance by removal of antibiotics from use was an idea that was put forth. Generally mutation in a bacterial gene to confer resistance often results in a loss of fitness leading to slower growth. Therefore it was believed that removal of antibiotics would cause selection against these bacteria when competing with non-resistant, fit bacteria. Unfortunately, for the most part, this has been shown to be ineffective. Bacteria often acquire compensatory mutations that allow them to grow as fast as or even faster than wild type, non-resistant bacteria. Consequently, the selective advantage of the non-resistant bacteria and the selection against the resistance mutation is eliminated.
The discovery of antibiotics was a leap in modern medicine. The work on penicillin by Fleming, Chain and Florey was followed by many scientists and pharmaceutical companies in search of new antibiotics with different modes of actions. They have been able to stop the growth or kill many different kinds of microorganisms. However, the bacteria in particular have proven to be much more innovative and adaptive than scientists had imagined. Bad practices and mismanagement have only exacerbated the situation. We could soon return to a state of medical health that was as dire as that which occurred prior to antibiotic use. However, with more research, education of the public, and well thought out government regulations, we will be well on our way to slowing down the resistance phenomenon.
References
1. Fleming A. (1946). Chemotherapy: yesterday, today and tomorrow. In (Hunter P.A., Darby G.K., Russell N.J. eds) Fifty Years of Antimicrobials: Past Perspectives and Future Trends. 53rd Symposium of the Society for General Microbiology. pg. 1-18. Cambridge University Press.
2. Ericsson H. (1981). Glimpses from 40 years of clinical bacteriology at Karolinska Hospital. In EAL Course Compendium 2002. p.4-19. EAL Solna, Sweden
3. Davies J. (1994). Inactivation of antibiotics and dissemination of resistance genes. Science 264:375-82.
4. Zahner H., Fiedler H. (1995). The need for new antibiotics: possible ways forward. In (Hunter P.A., Darby G.K., Russell N.J. eds) Fifty Years of Antimicrobials: Past Perspectives and Future Trends. 53rd Symposium of the Society for General Microbiology. p.67-84. Cambridge University Press.
5. Cunha B. (1998). Antibiotic resistance: control strategies. Critical Care Clinics 14(2):309-27.
6. Rowe-Magnus D.A., Guerout A.M., Ploncard P., Dychinco B., Davies J., Mazel D. (2001). The evolutionary history of chromosomal super-integrons provides an ancestry for multiresistant integrons. Proceedings of the National Academy of Sciences USA 98:652-7.
7. Barbosa T.M., Levy S. (2001). Antibiotic use and resistance: what lies beneath! In APUA Newsletter. Vol 19 No. 1. pgs. 1-3
8. Finkelstein J., Stille C., Nordin J. Overuse of antibioitics in children decreasing in United States. General Pediatrics IV Platform Session. Presented at the Pediatric Academic Societies Meeting. May 2002. Baltimore.
9. Rowe-Magnus D.A., Davies J., Mazel D. (2002). Impact of integrons and transposons on the evolution of resistance and virulence. In (Hacker, J & Kaper, J.B. eds.) Pathogenicity Islands and the Evolution of Pathogenic Microbes Vol 2. p.167-188. Springer-Verlag.
(Art by Fan Sozzi)