IDENTIFICATION AND LOCALIZATION OF THE SANEW GENE IN MAGICAL WOLFSBANE (ACONITUM ALUPUS)
Annals of Praetachoral Mechanics (2016). Vol 2. Advanced online publication. download pdf
ABSTRACT
For more than 100 years, those suffering from werewolfism have relied on the Wolfsbane Potion to maintain cognitive abilities during transformation. Recent developments in biotechnology and biochemistry have allowed for in-depth study of the chemistry and genetics of magical plants such as Aconitum alupus, the most important herbal component of Wolfsbane Potion. A recent study identified mesaconitine as the potion-active metabolite of A. alupus, though the details of its biosynthesis have not yet been elucidated. In this paper we reported the discovery of the gene SANEW which was identified using Ilumina sequencing to compare the transcriptomes of wild type A. alupus and a mutant non-magical A. alupus. We proposed this gene encodes a DOXP synthase involved in the synthesis of mesaconitine. To further verify the involvement of this gene in the synthesis of mesaconitine, we produced knockout A. alupus lines using an RNAi hairpin construct. Relative expression levels of the SANEW gene in these knockout lines measured using quantitative RT-PCR were found to be between 10% and 15% compared to wild type. Furthermore, levels of mesaconitine were measured in the knockdown mutants using High Performance Liquid Chromatography, and found to have a reduction in mesaconitine content 1.7-7.9% of wild type. Low expression levels of the SANEW gene correlated with low levels of mesaconitine, which supported the hypothesis presented regarding the role of SANEW. These findings take us a step closer to being able to produce mesaconitine synthetically and ultimately develop a drug therapy as an alternative to use of the Wolfsbane Potion, which is not presently accessible to all patients affected by werewolfism due to the high level of skill and experience required to brew it.
Keywords: SANEW, Aconitum alupus, Lycanthropy, Transcriptome, Mesaconitine synthesis, HPLC, Potion-active compounds, RNAi knockdown, methylerythritol phosphate pathway
Introduction
Magical Wolfsbane is an important magical plant belonging to the family Ranunculaceae. The genus Aconitum contains approximately 250 species, but the most scientifically important of these is Aconitum alupus, a perennial commonly found in northern Europe (Spore, 1408). A. alupus is of great interest to Healers, potioneers, and researchers due to its unique medicinal properties. Paramount of course is its use in the Wolfsbane potion developed by Damocles Belby (1876), which mitigates some of the effects of lycanthropy, more commonly known as werewolfism. Lycanthropy is a condition that has plagued wizards and muggles alike for centuries (Borelli et al., 1975). This condition is controlled by lunar patterns, where an affected person undergoes a transformation into a vicious and deadly wolf-like creature for the duration of every full moon. Whilst transformed, the werewolf becomes non-sentient and loses the cognitive ability associated with their human self (Lockhart, 1986). A transformed werewolf is neither aware of his or her identity, nor the identities of those around them, and will kill any human, even friends and family, given the opportunity. When used properly as a treatment, the Wolfsbane Potion allows the affected person to hold onto his or her humanity during their time as a werewolf, preventing damage to property and people. However, the potion does not prevent the transformation itself. The potion is very difficult to brew, one reason being that A. alupus contains a toxin called aconitine in addition to the potion-active compound; in fact, when brewed incorrectly, the potion is lethal to the drinker (Jigger, 1990). Thus, the development of improved treatment options for those affected by lycanthropy is of great interest to the wizarding community.
The phytoconstituants of A. alupus have been evaluated to elucidate the metabolites responsible for the plant’s magical qualities. Lupin and Weasley have reported mesaconitine, an alkaloid similar to aconitine, as the potion-active metabolite which, like aconitine, is synthesized in the methylerythritol phosphate pathway (MEP) (Lupin & Weasley, 2014). Recently, it was discovered that the A. alupus collected on the grounds of Hogwarts School, resulted in an ineffective Wolfsbane potion when brewed according to the original Belby method (1876). After further inspection by Lupin and Weasley, it was discovered that these plants were mutants which contained no mesaconitine. This mutation could be an accidental result of the large amount of spell casting which is practiced at the site of collection. The resulting loss-of-function phenotype identified in these mutants suggests there may be a direct linkage between a gene involved in the MEP, and that identification of this gene could further our understanding of mesaconitine synthesis in A. alupus. In this study, we aimed to identify the function of this gene using transcriptome analysis between mutant and wild-type A. alupus. In addition, we wish to replicate the loss-of-function phenotype using RNAi-knockdown of the identified candidate gene to provide further insight into the functional role of this gene in mesaconitine synthesis in A. alupus.
Materials and Methods
Transcriptomics
RNA Extraction and cDNA synthesis
Mature leaf tissue was collected from 6-week-old A. alupus plantlets, flash frozen using liquid nitrogen, and stored at -80°c until needed. The SV Total RNA Isolation System (Promega Corporation, USA) was used to extract RNA and column purified as specified by the manufacturer’s instructions. The quality of the RNA was tested using an Agilent 2100 Bioanalyzer RNA 6000 NANO assay (Agilent Technologies, USA). Starting with 1 µg of RNA, complementary DNA (cDNA) was constructed using TruSeq RNA Sample preparation protocol v.2 instructions (Low Throughput protocol, Ilumina Inc.) The quality of the RNA was tested using a NanoDrop 2000 UV-Vis Spectrophotometer and samples with a 260 nm/280 nm ratio between 1.8–2.1 were used to synthesize cDNA. Using the The TruSeq RNA Sample Prep Kit v2 (Illumina Inc., USA) Total RNA samples (1µg) were reverse-transcribed into double-stranded cDNA. The cDNA was stored at -20°c until needed.
Sequencing and processing of RNAseq data
Sequencing was performed on the Illumina HiSeq 2000 paired-end at 2 x 101 bp using the TruSeq SBS Kit v3-HS (Illumina Inc., USA) The reads were aligned to the A. alupus genome and transcriptome with Tophat v 1.3.3. The analysis of Tophat files was performed on the CLC Genomics Workbench v. 5.5.1 (CLC bio, Denmark), following the manufacturer’s instructions. The RNA sequencing (RNA-seq) data was mapped against the unannotated A. alupus genes. Differentially expressed genes were found using the CLC Genomic Workbench. Only statistically significant differentially expressed genes were considered (p < 0.01). For gene ontology functional annotation, the differentially expressed genes were loaded on Blast2Go v.2.5.1, a tool for functional annotation of (novel) sequences to annotate for biological processes, and molecular functions. The webtool Mapman v.3.5.1 (http://mapman.gabipd.org/web/guest) was used to visualize in which pathways the differentially expressed genes are categorized according to their annotations.
Knockout Transgenics
Construct development
RNAi-mediated transcript down-regulation was used to further investigate the function of SANEW, with the objective of suppressing the endogenous candidate gene SANEW in wild type A. alupus to create knockout plants which do not produce mesaconitine. A ≈ 400 bp hairpin RNAi-SANEW construct was generated using the pHannibal7 vector, and transformed into wild type A. alupus. The partial sequence fragment was obtained from A. alupus cDNA. The sequence was amplified by PCR with BamHI and Xhol restriction enzyme sites using PCR primers (5’-TGGATCCGCATTGCTATTGGTACGTT-3’ and 5’-TAAGCTCGAGTACAATGCGGCTAGT-3’) in order to ensure the directional cloning into pHannibal7 vector. Another set of primers (5’- TTAGCATGCACGTGTACCGTAC-3’ and 5’-GGATCCGAGACTATTGACAG-3’) containing BbuI and KpnI restriction enzyme sites was used to clone the same fragment of SANEW sequence in the reverse direction to facilitate the later formation of the RNA hairpin.
Agrobacterium-mediated transformation
A. alupus was transformed using Agrobacterium tumefaciens strain EHC211 containing the RNAi construct using standard leaf disk inoculation technique. The bacteria were incubated in liquid woody plant medium with 2.5% sucrose and 100 μM acetosyringone for 12 hours. Leaf disks were cut from leaves of wild type A. alupus grown in tissue culture, and after cocultivation with the bacteria for 1 hour, plated onto solid woody plant medium with 3.5% (wt/vol) agar, 1.3% (wt/vol) phytagel and 0.08% μM each α-naphthalene acetic acid, thiodiazurone and 6-benzylaminopurine (medium1). After 3 days, we transferred leaf disks to medium2 which is identical to the last with the addition of cefotaxime sodium salt (250mg/L) and carbenicillin disodium (500mg/L) for another 3 days to kill the agrobacterium. The leaf disks were then transferred to medium2 with added kanamycin (25mg/L) in order to select for positive transformants. After 10 weeks, isolated shoot tips were transferred to woody plant medium without antibiotics and left to grow in tissue culture.
Transgenic Plant Confirmation
We checked for positive transformants by PCR amplification with a vector-specific forwards promoter (5’-GCAGCTGACGCGTACACAACAAG-3’), and SANEW sequence-specific reverse promoter (5’-CCACCAATGCAAGCAGTCAGCGCAT-3’). The transcription levels of the SANEW gene were checked in the transformants that were deemed positive from the PCR genotyping using quantitative RT-PCR. Quantitative RT-PCR reactions consisted of 10µl SsoFast EvaGreen Supermix (Bio-Rad, USA), 5 pmol primers for SANEW (forward 5’-CCGTGCTGATGCAGGCATG-3’ and reverse 5’-CGGATTACACTGACTGCGAAG-3’ primers) or TIF5A genes, 1µl cDNA (1:5 dilution), and water to a total volume of 20µl. RT-PCR was performed using a CFX96 Real-Time PCR Detection System (Bio-Rad). The program run was one cycle at 95°C for 30 sec, 40 cycles at 95°C for 5 sec, 59°c for 5 sec, followed by once cycle of 55°c to 90°c to calculate the melting curve. TIF5A was used as a reference gene to calculate relative expression.
Plant Growth Conditions
Seeds were collected from transformants with low level transcription of the SANEW gene, and were planted in April 2014 following methods provided by Butola and Badola (2008). Seeds were washed with double distilled water 3 times, suspended in water and then sown into 10 35cm3 pots with approximately 20 seeds per pot. A 1:1 mixture of manure and sandy loam soil was used. All pots were placed in a greenhouse with a temperature between 21°C and 25°C and RH between 45.5-91.0%. After 3 weeks of growth, seedlings were thinned to the strongest seedlings, leaving 3 plants per pot.
High performance liquid chromatography
After transcript levels were measured, high performance liquid chromatography (HPLC) was used to quantify the levels of the alkaloid mesaconitine in knockout A. alupus as described by Wang and Xu et al. (2005). We used an HPLC system with an XTerraTMRP24 column and 0.03 M ammonium hydrogen carbonate-acetonitrile as elution mobile phase. First we ran the system with a series of mesaconitine standard concentrations. This allowed us to calculate the regression equation by looking at the linear relationship of mesaconitine concentration and the area under the peak. This equation was used to identify mesaconitine in our samples by comparing the retention time and the UV spectra data from our samples to those of the standard.
Results and Discussion
Identification of candidate gene
The mutant A. alupus was sequenced using Illumina® sequencing. The RNAseq data was aligned to the A. alupus reference genome, and the genes were annotated with possible biological functions. Initially, there were 143 differentially expressed genes in the mutant A. alupus. Since mesaconitine is synthesized via the methylerythritol phosphate pathway (MEP), in order to narrow down which of these 143 genes could be of importance, we looked at the RNAseq data using Mapman to help visualize which differentially expressed genes were annotated as being involved in biosynthetic pathways of secondary metabolites (Fig. 1). After having been screened through Mapman, 31 genes were identified in the MEP. The most downregulated genes would be good candidates, which in Figure 1 are shown in shades of red. We looked at the annotation of the most downregulated genes, and three of them were annotated to have an SxNxxH motif, placing it in the Dxs family of DOXP synthases (Mandria et al., 2013). These catalyze reactions in the synthesis of metabolites similar to aconitine and mesoaconitine in other species such as Valerian (Valeriana officinalis) or Asphodel (Asphodelus ramosus) which are the main ingredients in the Draught of the Living Dead (a powerful sleeping potion) and the Draught of Peace (a potion relieving anxiety and agitation) respectively (Sparks et al., 2001; White and Pearson, 1998).
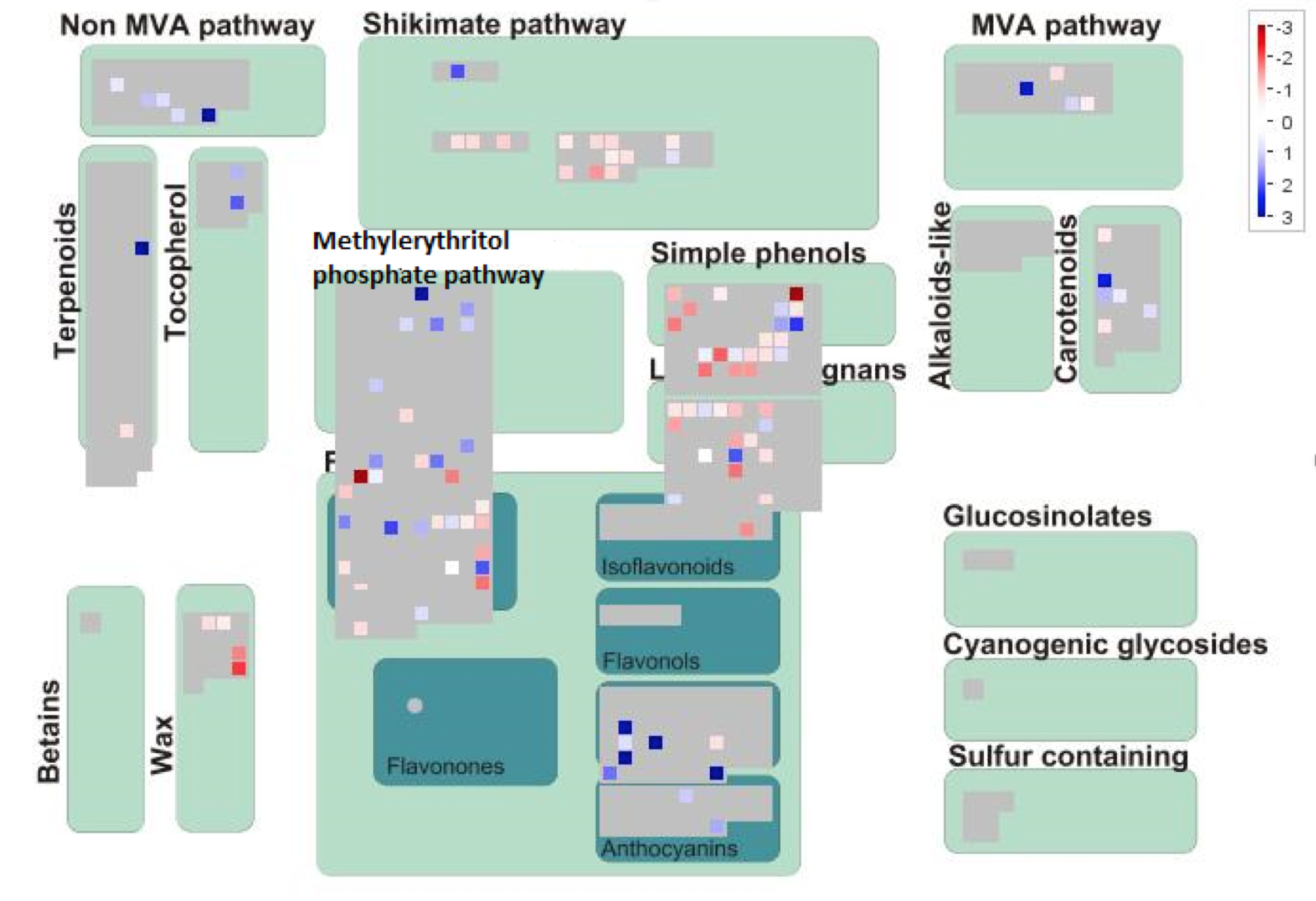
(CLICK ON IMAGE TO ENLARGE) Figure 1: Mapman Secondary Metabolite overview map of differentially expressed genes in mutant A. alupus. Down-regulated genes marked in shades of red, and up-regulated genes marked in shades of blue. Putative gene SANEW in the MEP marked with the red arrow.
Knockout Transgenics
Genotyping via PCR revealed 9 putative positive lines of RNAi-SANEW transformants. Quantitative RT-PCR showed that all 9 of the recovered lines had suppressed transcript levels of the SANEW gene (Fig. 2). Relative expression levels of the SANEW gene in these knockout lines measured using quantitative RT-PCR were found to be between 10 and 15% compared to wild type. Although the reduction in expression varied between lines, all lines showed a significant reduction in expression relative to wild type. The knockout A. alupus plants were viable, and did not show any visible phenotype to suggest changes in general physiology, but this should be researched in more depth in the future, to be sure.
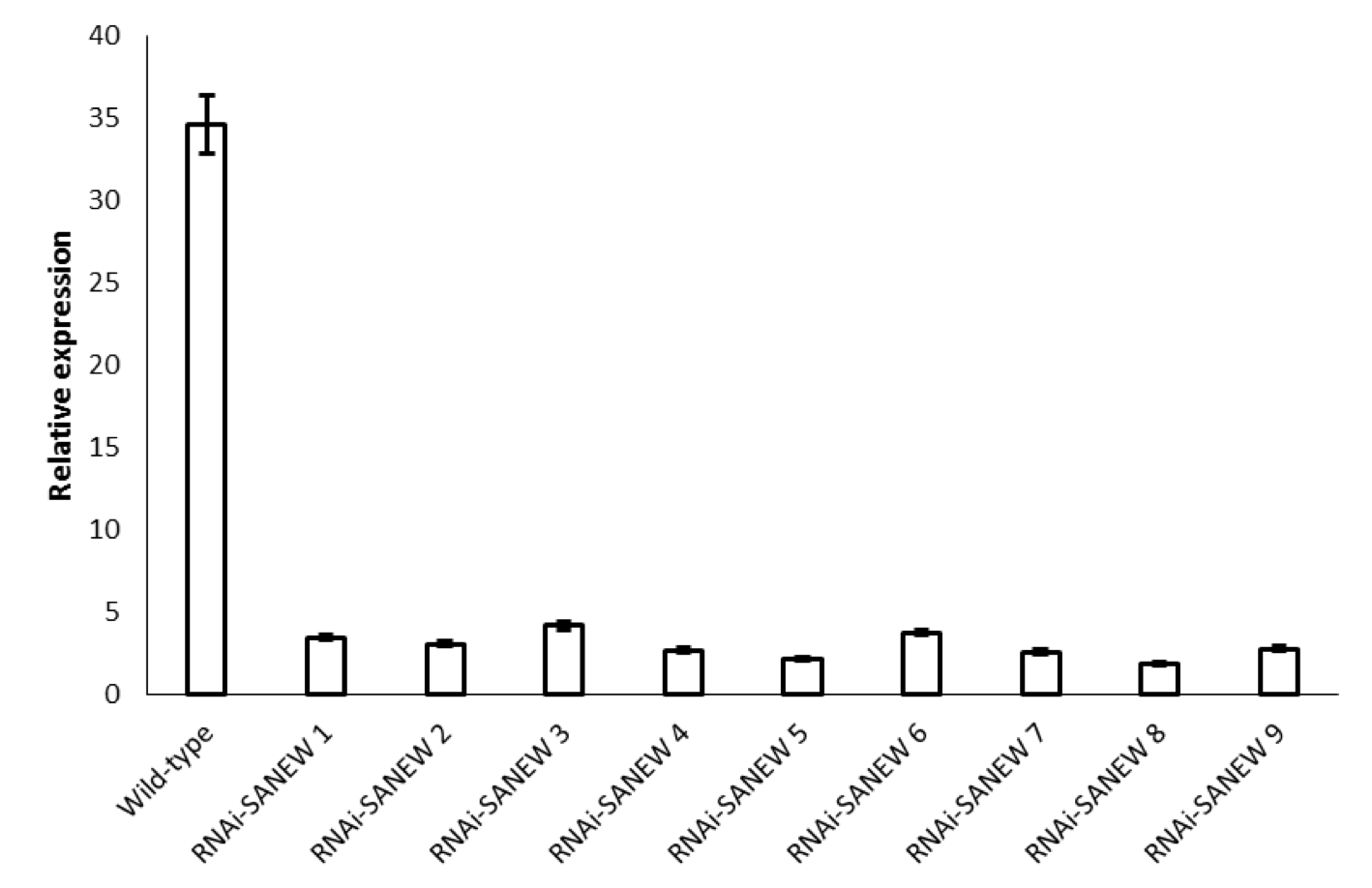
(CLICK IMAGE TO ENLARGE) Figure 2. Relative expression levels as measured by quantitative RT-PCR of RNAi-SANEW knockout A. alupus plants (n=3). The error bars correspond to standard error of the mean, based on three replicates.
High performance liquid chromatography
HPLC was used in order to quantify mesaconitine levels in both wild type and RNAi-SANEW A. alupus plants. The linear relationship of the standard concentrations to the peak area equation was as follows;
Y=8.87e+003x-1.48e+004
Where x = concentration of active ingredient (μg/ml), y = peak area. The correlation coefficient of the equation is 0.99, indicating a linear relationship, as expected from the standard. The retention time of mesaconitine, as shown from the standard (Fig. 3), is about 47 min (average for series concentration is 47.11 min). This is in agreement with retention times found in the literature (Wang and Xu, 2005).
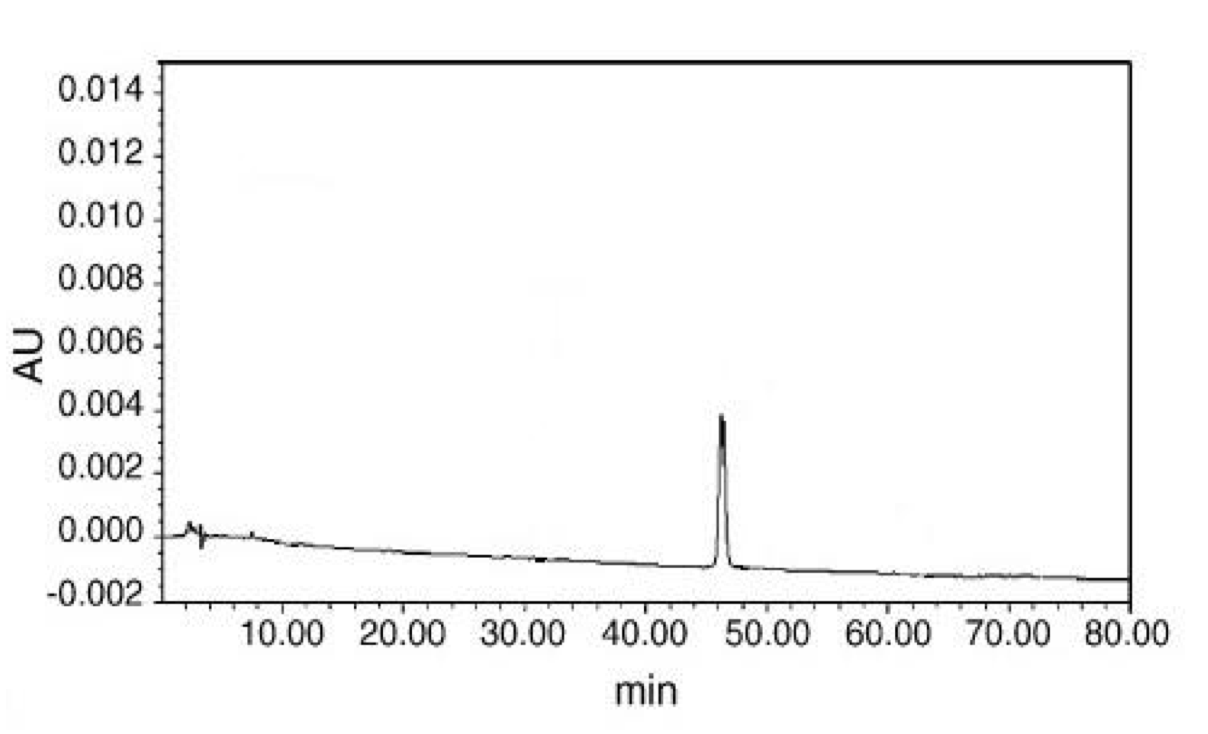
(CLICK ON IMAGE TO ENLARGE) Figure 3. Chromatograph of mesaconitine standards, showing the retention time of approximately 47 min, measured using an HPLC XTerraTMRP24 column and 0.03 M ammonium hydrogen carbonate-acetonitrile as elution mobile phase.
As expected, HPLC revealed that the RNAi-SANEW knockout plants had very low levels of mesaconitine relative to wild type levels. Wild type A. alupus showed a peak at 47 min, representative of the mesaconitine content (Fig. 4A), whereas the chromatographs of the RNAi-SANEW knockout plants did not have a peak correlating to mesaconitine (Fig. 4B, C, D). Also notice that, with the exception of the peak at 47 min, the RNAi-SANEW lines 4, 5, and 8 shared a similar chromatographic backgrounds to that of the wild type plant. This implies that the downregulation of SANEW had little impact on other aconitine-like compounds which have similar retention times (Wang and Xu, 2005)
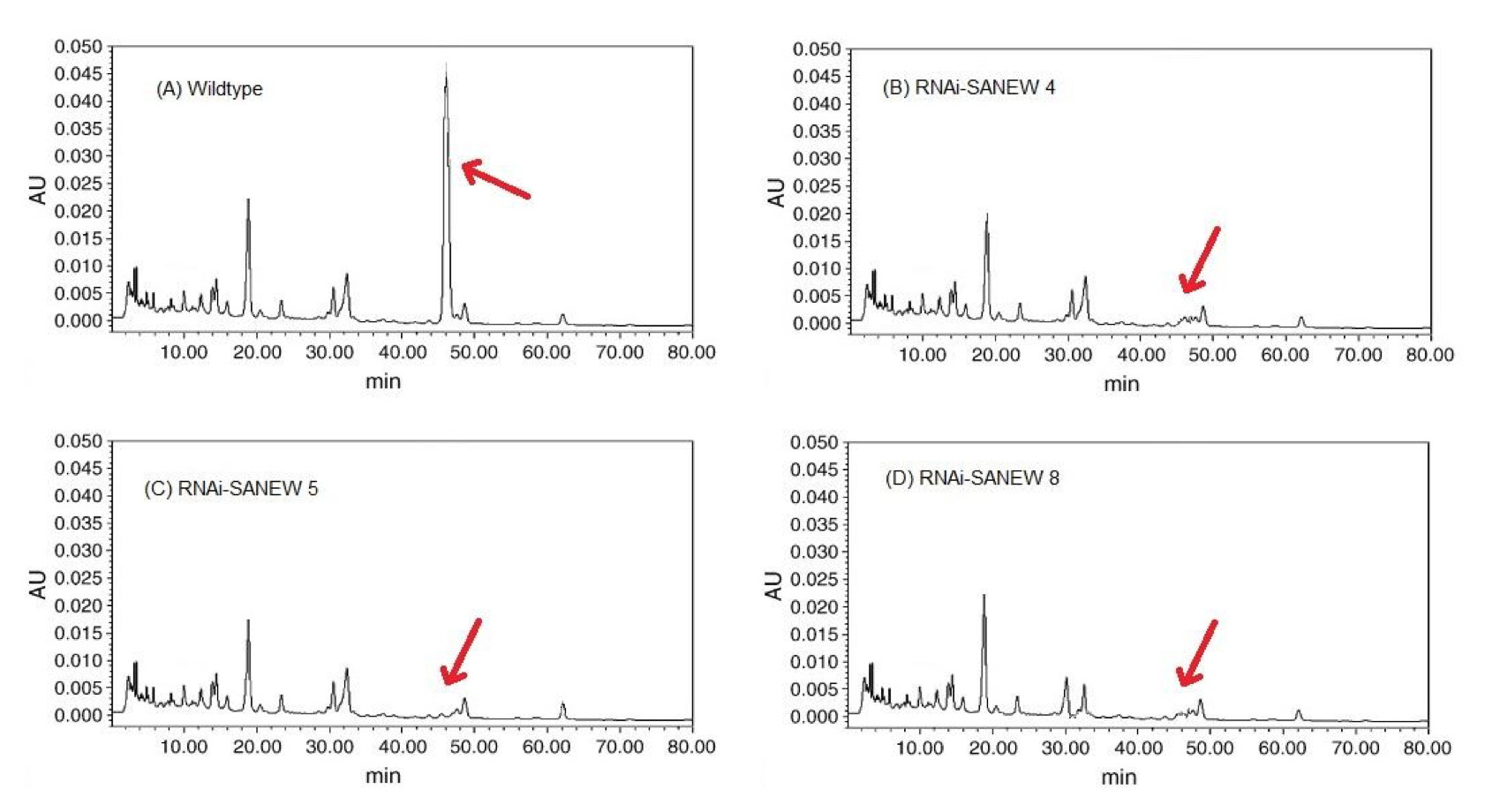
(CLICK ON IMAGE TO ENLARGE) Figure 4. A) Chromatograph of wildtype A. alupus showing peak at 47 min, corresponding to the retention time of mesaconitine. B, C, D) Chromatograph of A. alupus RNAi-SANEW knockout lines 4, 5 and 8 lacking a prominent peak at 47 min. corresponding to low levels of mesaconitine.
We used the regression equation from the standards and the peak areas obtained from the chromatographs of the separate lines to calculate mesaconitine content (Table 1). Calculation results are consistent with the outcome of transcript abundance levels. As expected, plants with low levels of expression of SANEW gene also had low mesaconitine levels.
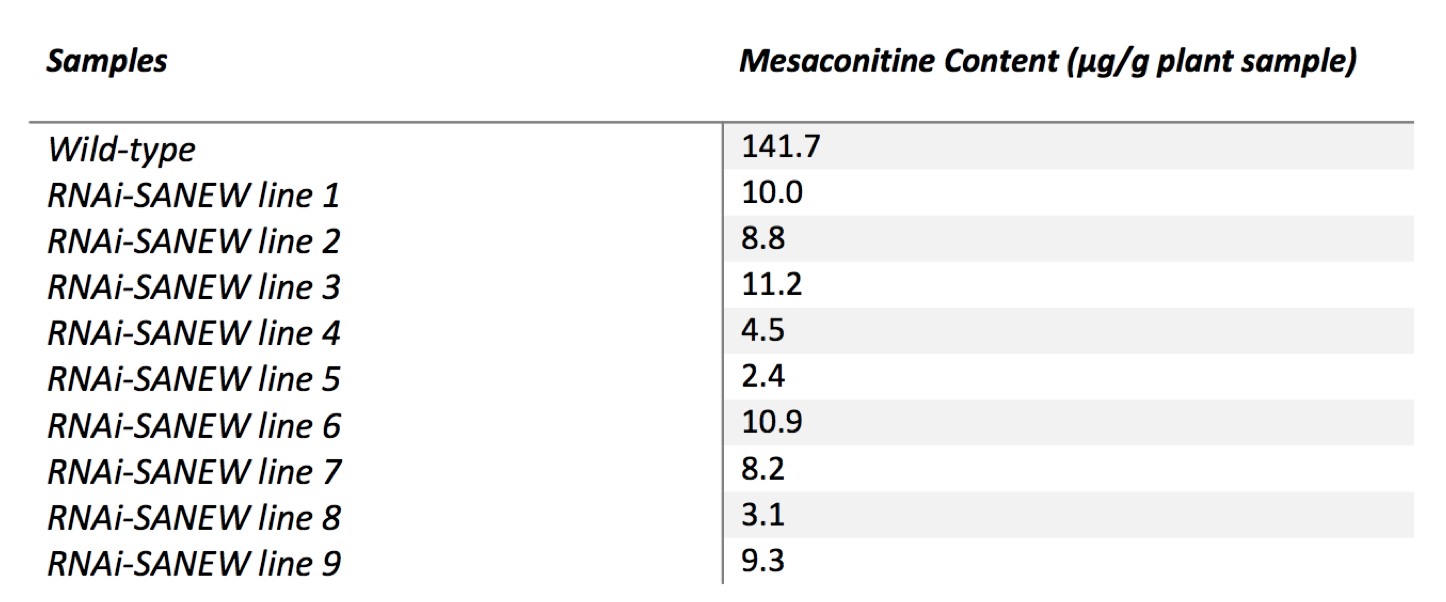
(CLICK ON TABLE TO ENLARGE) Table 1. Mesaconitine content (μg/g plant sample) of the wild type and RNAi-SANEW knockout A. alupus. (n=5)
Conclusions
For over 100 years, Magical Wolfsbane has been a vital part of the lives of those affected with lycanthropy. It is the main ingredient in the Wolfsbane Potion, and as a result has been studied for decades – although the in-depth study of its molecular biology and genetics has only more recently become possible. Next-generation sequencing techniques are allowing researchers to make great progress in elucidating gene function and biosynthetic processes of magical and medicinal plants. With the rapid evolution of next-generation sequencing techniques, increased sequencing precision, and decreased costs, these techniques are becoming accessible to all witch and wizard researchers.
Recently, work has been done on the A. alupus by Lupin and Weasley (2014) regarding the potion-active alkaloid mesaconitine, but this paper is the first to report on a putative gene responsible for the synthesis of mesaconitine. Comparative transcriptomics resulted in the identification of a putative gene that we have named SANEW. The results of our RNAi-mediated down-regulation of SANEW indicated that this gene appears to be important to the synthesis of mesaconitine. We hypothesize that the SANEW gene encodes a DOXP synthase as it has an SxNxxH motif characteristic of the Dxs family of enzymes. These are known to be involved in the synthesis of mesaconitine-like alkaloids.
Future studies introducing the SANEW gene into non-magical plants, such as Arabidopsis thaliana, are proposed to see if this would induce the production of mesaconitine, would allow for further confirmation that the SANEW gene is required for its synthesis. Other future research of the SANEW gene should include the isolation and characterization of the DOXP synthase, followed by kinetic analysis of the enzyme. Through the continued study of this enzyme we hope find a way to synthesize mesaconitine in the lab without dependence on A. alupus as a source material. The proper brewing of the Wolfsbane Potion requires a level of experience and skill in potion making that very few Potion Masters have. If improperly brewed, the potion is in fact lethal to the drinker. Consequently the applications of a synthetically produced mesaconitine as a substitution to the Wolfsbane Potion, if effective, would be of enormous value. This would allow access to treatment for those afflicted with lycanthropy without the need for a Potion Master, and scale of production could be increased.
References
Borrelli S, Braida J, Cassileth T, Motti M (1975) Lycanthropy: a history. Behavioural Studies and Analysis 25: 45-66
Butola JS, Badola HK (2008) Propagation conditions for mass multiplication of three threatened Himalayan high value medicinal herbs. International Plant Genetic Resource Newsletter 153: 43-47
Jigger A (1990) Advanced Magical Drafts and Potions, Tenth Edition. Magitext Inc,
London, UK, 999 p.
Lockhart G (1986) Wanderings with Werewolves. Magitext Inc, London, UK, 457 p.
Lupin E, Weasley R (2014) Potion-active metabolite mesaconitine; Isolation and Identification in Aconitum alupus. Journal of Plants with Magical Uses 131: 213-221
Mandria R, Penova S, Lining M (2013) The Dxs family of DOXP synthases; A Review. Annual Reviews 204: 467-490
Sparks A, Welsh M, Fan Q (2001) Chemical analysis of a traditional herb Valeriana officinalis frequently used in potions. Biochemical Journal 155: 747-759
Spore P (1408) One Thousand Magical Herbs and Fungi. Magic Associates, London, UK, 893 p.
Wang X, Xu B (2005) Improved quantitative determination method of diterpenoid alkaloids in four species of Aconitum by high performance liquid chromatography. Journal of Pharmaceutical and Biomedical Analysis 68: 1031-1034
White P, Pearson D (1998) Anti-anxiety effects of the herb Asphodelus ramosus and the determination of the active ingredient. Journal of Biomedical Science 171:77-90
