THE EFFECT OF NOVEL TELOMERASE REVERSE-TRANSCRIPTASE wTERT2 ON THE PATHOGENESIS OF WOOKIEE IDIOPATHIC PULMONARY FIBROSIS
Annals of Praetachoral Mechanics (2015). Vol 1. pp81 download pdf
ABSTRACT
The Wookiee population on Kashyyyk is responsible for a majority of the labour workforce in the Mid Rim. Maintaining and improving Wookiee lifespan and productivity is vital for Imperial operations in the area. The long Wookiee natural lifespan of 400 standard galactic years is truncated in approximately 3% of the popula- tion. These Wookiees are afflicted with Idiopathic Pulmonary Fibrosis (IPF) and show early-onset leukocyte se- nescence. Once diagnosed, Wookiees live for an average of 30 more years before death. This translates into a 70% reduction of life for affected Wookiees, and there is no cure. This paper describes a novel genetic effector for premature aging and IPF in Wookiees in the form of a previously uncharacterized gene, wTERT2. A high- throughput gene profile led to the discovery this highly upregulated gene after 80 years of age (104.3 fold change, p<0.0001). A transgenic rodent model using the Kashyyykan Ikov was constructed, replacing the wildtype iTERT with wTERT and wTERT2. The rodents expressing wTERT2 exhibited 24% longer lifespan (p<0.05) as well as abrogated age-related comorbidities such as shorter leukocyte telomere length (LTL, p<0.001), decreased bone density (p<0.002), loss of subcutaneous adipose (p<0.04), and glucose intolerance (p<0.01).
Performing targeted exome sequencing on the wTERT2 locus in the Wookiee cohort revealed a strong asso- ciation (p<0.001) between IPF and a A2196X single nucleotide substitution. The mutation resulted in the substitu- tion of Threonine to Alanine or Serine in the 732nd residue. Due to the low penetrance of the IPF phenotype in the Wookiee population, it is thought that the mutation results in a haploinsufficiency in which mutated wTERT2 is insufficient for normal phenotype.
This study introduces a new focus in the field of Wookiee aging. The concept of a second, more aggressive telomerase reverse transcriptase is not seen in other species in the galaxy. Further research into wTERT2 could yield solutions to accelerated aging and IPF in Wookiees. The result could significantly improve the productivity of the Wookiee labour force, which would have a dramatic impact on Imperial operations in the Mid Rim.
Key words: wTERT2, Wookiee, Idiopathic Pulmonary Fibrosis
Introduction
The Imperial Health Research Initiative (IHRI) seeks to improve health and longevity for all species in the Galactic Empire. The Wookiees (Wookiee wookiee) are an indentured species native to Kashyyyk, a planet consisting of most of the Mid Rim labour force. Maintaining health and longevity of the Wookiee population on Kashyyyk is crucial to the welfare of large-scale military projects such as the Maw Installation, which in turn is essential for the security and viability of the Empire.
The Wookiee is a species of large humanoids characterized by their superhuman strength, full body of hair, and longevity. The natural lifespan of a Wookiee is approximately 400 standard galactic years. Although the Wookiee culture recognizes adulthood at the age of 18, it has been shown that the average Wookiee reaches physiological maturity at the age of 80. How- ever, roughly 3% of the Wookiee population present with early-onset leukocyte senescence, and are at high risk for Idiopathic Pulmonary Fibrosis (IPF). IPF is characteristic of telomerase-related genetic disorders in other galactic mammal species. In many such species, telomerase is a highly conserved enzyme that functions to preserve telomere length and attenuate midi-chlorian oxidative damage, which are both strong biomarkers for cellular aging. Mutations in essential telomerase genes, TERT and TR, have been implicated in pulmonary fibrosis in 86% of species studied, although no such causal genetic defect has been found in Wookiees thus far. Aggregate planetary clinical data show that 81% of all IPF diagnoses in Wookiees occur when the patient is between the ages of 80 and 100, similar to the age at which Wookiees reach full maturity. Positive family history is the most prevalent risk factor, with up to 77% of Wookiee IPF patients reporting affected family members. Reduced leukocyte telomere lengths have also been observed in age-matched studies of Wookiees and other galactic mammals with IPF. Similar to other galactic mammals, Wookiees express telomerase reverse transcriptase (wTERT) in pluripotent stem cells during development, with no wTERT activity detected in most somatic cells as an adult. This suggests a physiological link between the specific age at which Wookiees reach maturity and an unknown genetic effector of age and cellular senescence.
This paper hypothesizes that the physiological change that occurs with maturity at around 80 years of age in Wookiees is accompanied by transcriptional activation or deactivation of a previously unknown genetic effector of age and cellular senescence. Additionally, we believe that a genetic mutation in this unknown effector is associated with accelerated aging and IPF in Wookiees. By performing exome gene expression profiling on 646 Wookiees both with and without IPF as well as over and under the critical age of 80, we revealed several candidates for putative age-related proteins, and explored their potential functional roles in prematurely aged Wookiees.
Results
Microarray Reveals Putative Age-Related wTERT Homologue
DNA microarray was used to generate expression profiles of prematurely aging, IPF-suffering Wookiees under the age of 80 (N=134) compared to a cohort of prematurely aging, IPF-suffering Wookiees greater than 80 years old (N=163). A cohort of normal Wookiees (N=151 and N=198, <80 and >80 respectively) were profiled as well. In both age groups, no gene expression differences reached significance when the normal and prematurely aged Wookiee cohorts were compared. The most significant hits in the gene array were between <80 year old compared to >80 year old Wookiees, regardless of IPF status. The top hit was a previously sequenced gene at the Reverse Transcriptase-Like Protein 4 (RTL4) locus on Chromosome 7 (of 13), P<0.0001 (Table 1). The RTL4 gene was expressed 104.3-fold higher in our microarray of the post 80 cohort, P<0.0001. The genetic sequences of the top hits with p-values <0.005 (Table 1) were queried through Basic Local Alignment Search Tool (BLAST) against the Wookiee wookiee Consensus Genome. Interestingly, the BLAST revealed that the RTL4 sequence has a Reverse Transcriptase Domain highly homologous with uncharacterized genes RTL3, RTL2 and the highly characterized wTERT gene (Figure 1A). Upon closer examination of the RTL4 gene compared to the wTERT gene, we found that both shared a POLIII Association Domain, and a protein BLAST revealed similar DNA binding domain residues (Figure 1B). Due to this striking similarity in functional residues and domains RTL4 will henceforth be referred to as wTERT2.
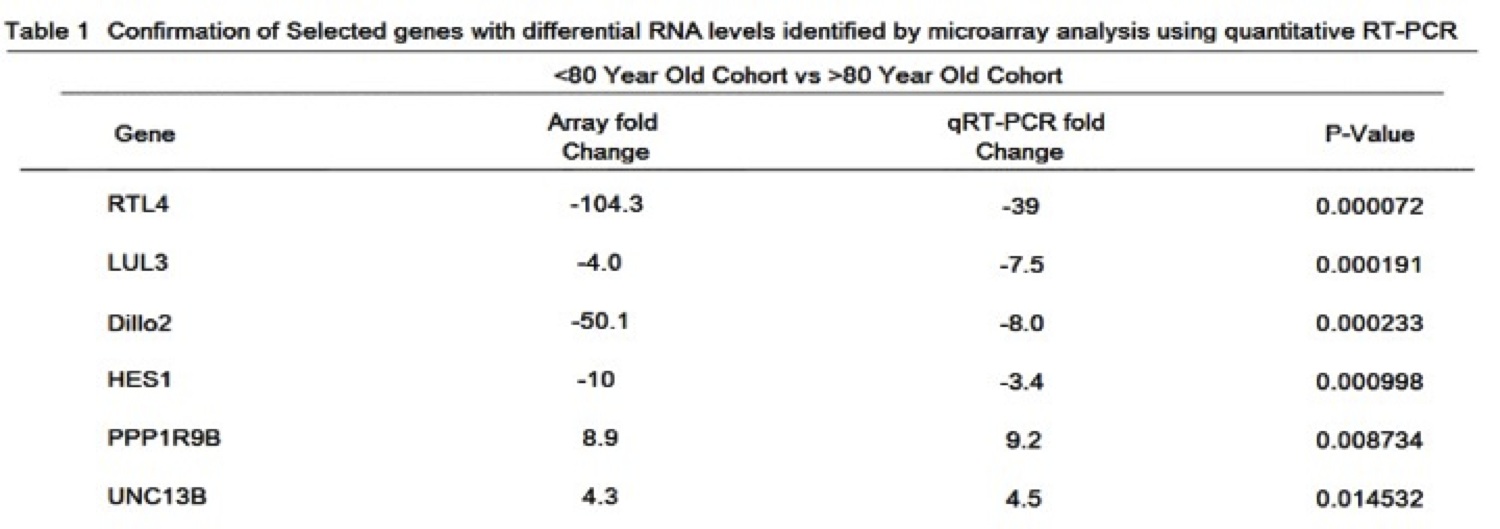
Table 1 (Click to Enlarge): Confirmation of selected genes with differential RNA levels identified by microarray analysis using quantitative RT-PCR.
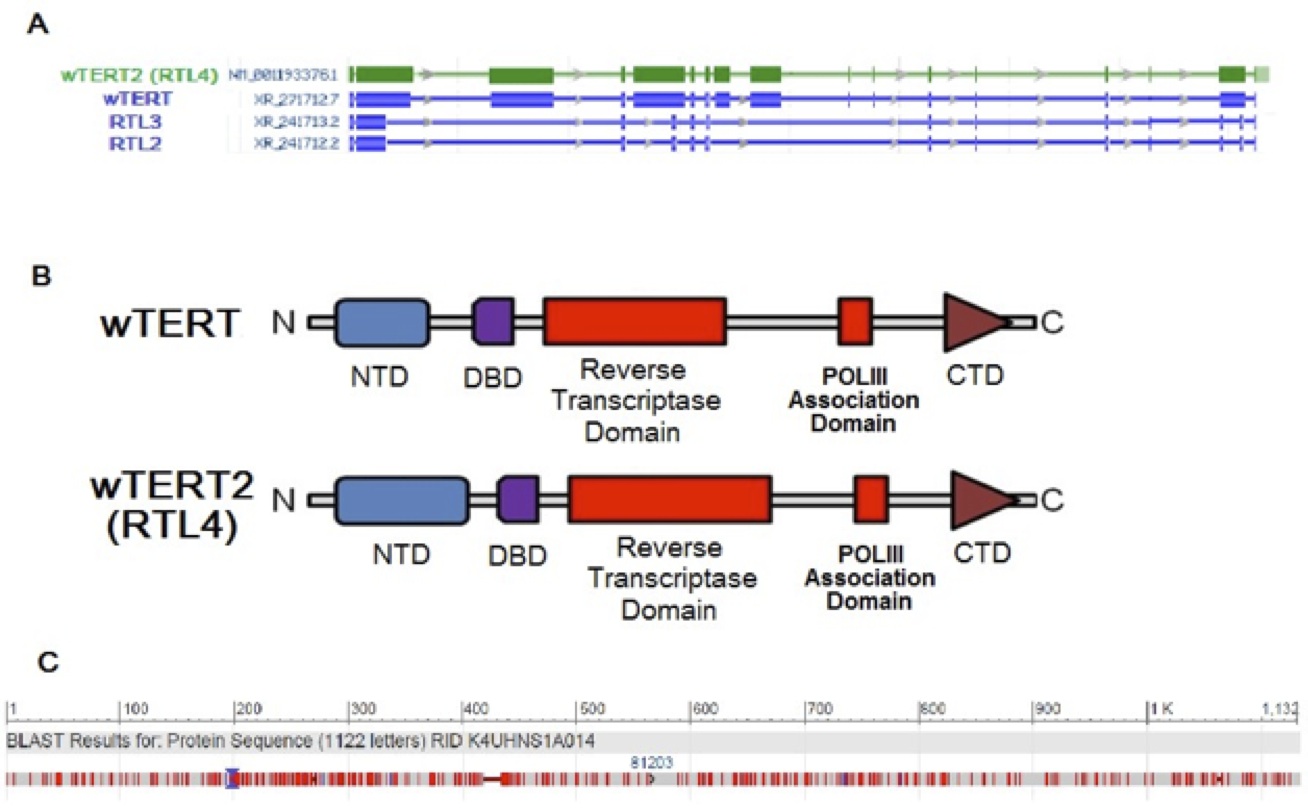
Figure 1 (Click to Enlarge): Genetic and Amino Acid Sequence and structural similarities between wTERT and Putative wTERT2.
A. Amino acid sequence similarities between top microarray hit wTERT2 (RTL4) compared to top BLAST hits.
B. Functional domain comparison of wTERT and wTERT2 (RTL4).
C. Amino acid sequence overlay of wTERT and wTERT2, conveying 76% sequence identity.
Development of a Kashyyykan Rodent Model for wTERT2 In Vivo Studies
In the selection of a model system to explore the effects of wTERT and wTERT2 in mammalian animals we sought to mimic the biological environment of the Wookiee species as closely as feasibly possible, therefore a close mammalian relative of the Wookiee from Kashyyyk would be the ideal candidate. The major challenge in identifying such a species is the harsh nature of most Kashyyykan species that are closely related to the Wookiee. Exploring a database of phylogenetic data of Kashyyykan species (5) we identified a ro- dent, the Kashyyykan Ikov, as most suitable for laboratory study.
The Ikov is an inhabitant of the Wroshyr tree forest of Kashyyyk, and shares a common ancestor relatively recently with the Wookiee species (Figure 2A). Despite having finely serrated teeth, the natural scavenging behaviour of the Ikov tempers its aggressiveness, making the creature easier to manage in a scientific research setting. Importantly the rodent expresses a homologue of wTERT, iTERT, with 76% matching residues (Figure 1C), and high conservation of the Reverse Transcriptase Domain (6).
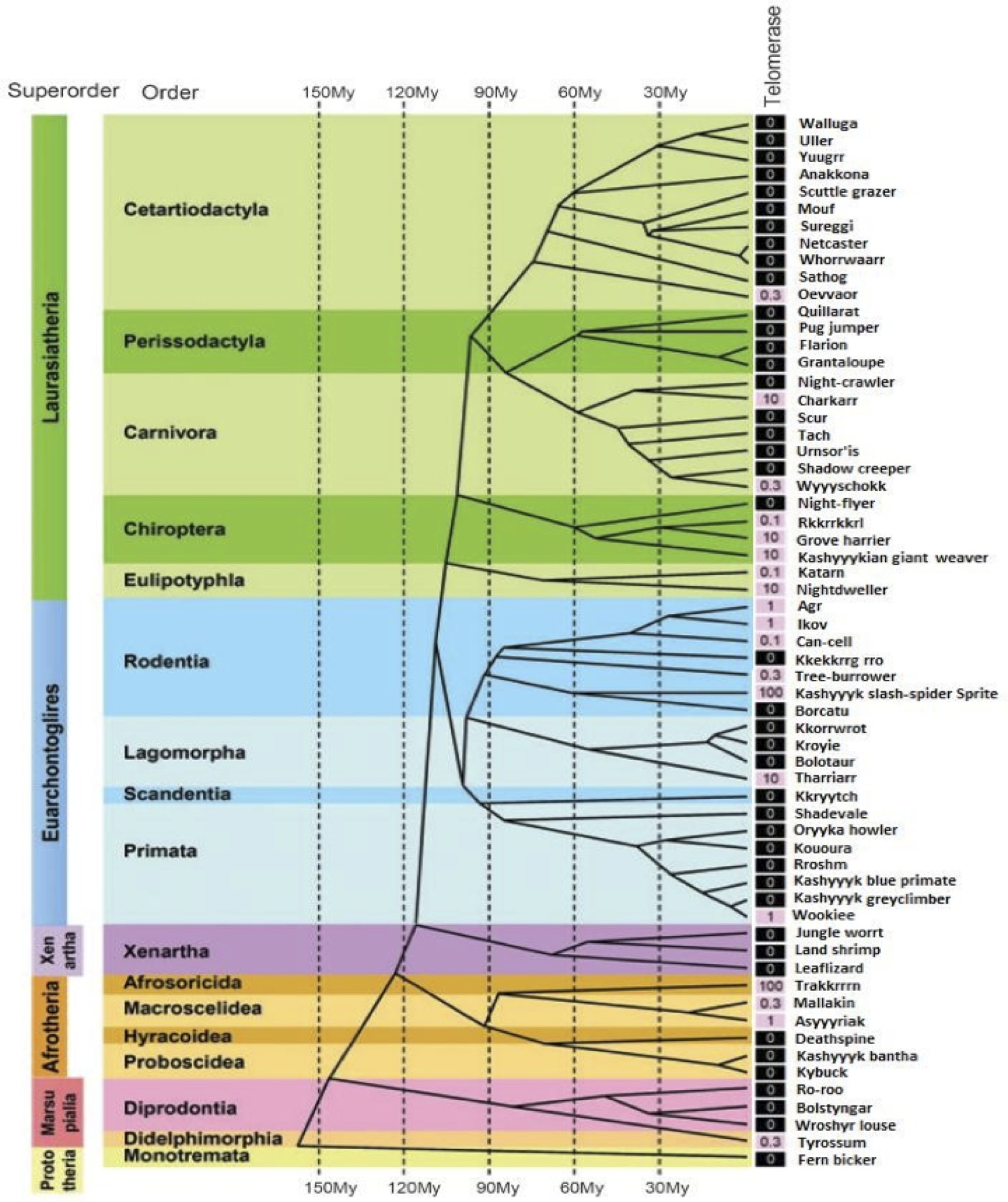
Figure 2 (Click to Enlarge): Evolutionary distribution of in the Kashyyykan mammal evolutionary tree (adapted from Gomes et al, 2011). Telomerase activity runs in evolutionary clades. Telomerase activity was detected with TRAP assay. Values are expressed as the percent- age of activity in reference to Ikov carcinoma cell line 4B4, shaded black if absent and pink if any activity was detected.
Employing Zinc-Finger Nucleases we created two novel transgenic Kashyyyk Ikov models, excising the wild-type iTERT gene and replacing the rodent’s iTERT with the GFP-tagged wTERT or GFP-tagged wTERT2 at the embryo stage (Figure 3A). Efficient inclusion of the wTERT and wTERT2 genes were confirmed by direct assessment of GFP fluorescence in living animals after birth and every six months thereafter (Figure 3B). Unexpectedly, using MMqPCR wTERT2 rodents were shown to have elongated telomeres (T/S Ratio) when compared to wTERT rodents (Figure 4A). These data were confirmed by flow-FISH on a subset of our cohort.
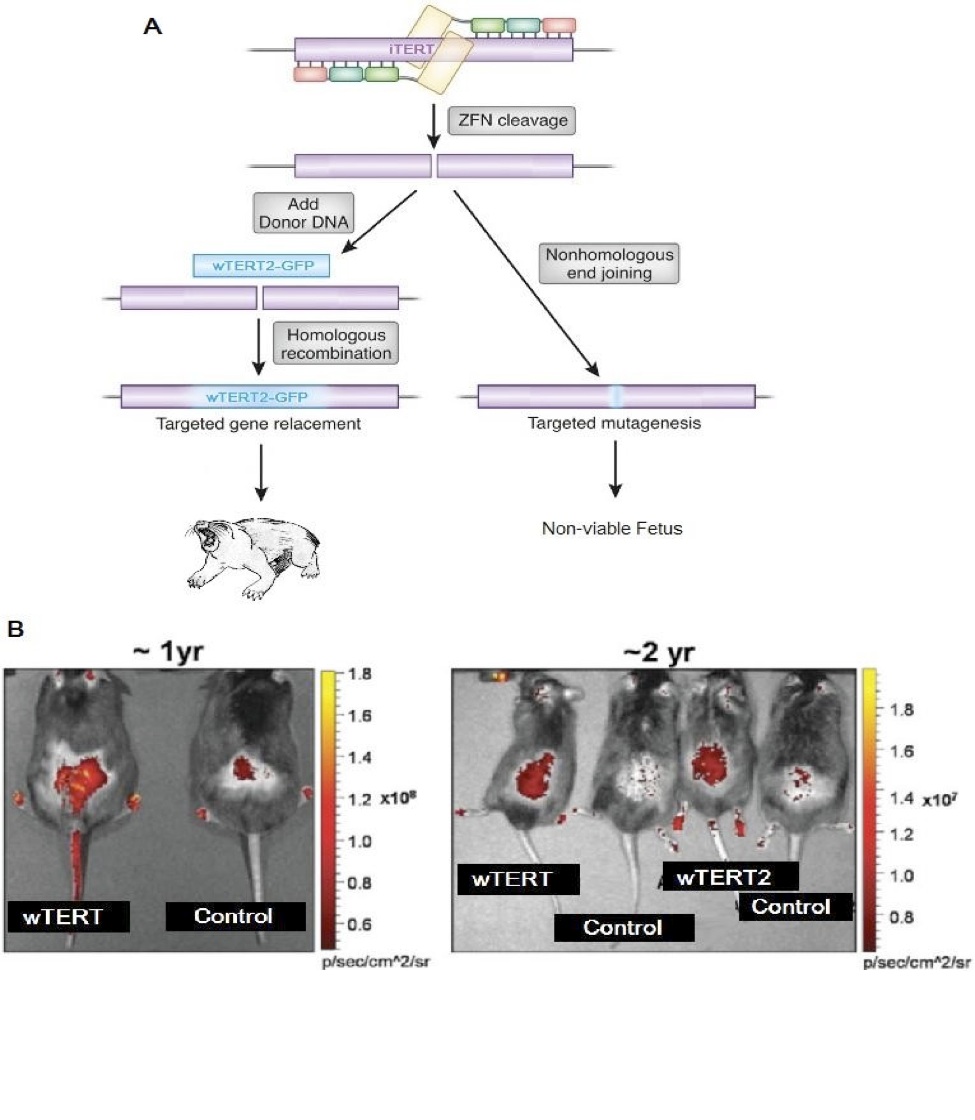
Figure 3 (Click to Enlarge): Development of a transgenic Ikov rodent model for the ectopic expression of wTERT and wTERT2 (adapted from de Jesus et al, 2000)
A. A zinc finger nuclease strategy was employed to genetic alter the developing embryo, excising the endogenous iTERT gene and replacing it with either a GFP tagged wTERT or a GFP tagged wTERT2.
B. Direct GFP fluorescence in shaved back skin of mice treated with the indicated ZFN vectors.
Age-Related Comorbitities Are Attenuated in wTERT2-Expressing Transgenic Rodents
Bone loss is a well-characterized marker of aging in most species in the galaxy due to the resorption of bone after failure of osteoblast cells (6). Assessing the bone mineral density (BMD) of the femurs of the trans- genic rodents we found that wTERT2 rodents showed slight, but significant maintenance of bone density compared to wTERT and control (iTERT) rodents at one year, but large and highly significant bone retention after two years (Figure 3B). This beneficial effect of wTERT2 is coincidental with the greater expression of GFP in the bone matrices of older mice compared to control rodents but not wTERT rodents.
Another biomarker of aging is the loss of the subcutaneous adipose skin layer, which can lead to a number of age-related comorbidities (7). As expected, the thickness of the subcutaneous fat layer in 2 year-old rodents was markedly reduced for all three rodent cohorts (Figure 4C). However, the two year old wTERT2 rodents showed a significant preservation of the subcu- taneous adipose skin layer, and were comparable to one year old rodents (Figure 4C). Again, we detected great- er expression of GFP in the white adipose tissue of wTERT and wTERT2 rodents compared to control rodents.
We next examined glucose intolerance as a marker of the aging process in our transgenic rodent models, since telomerase deficiency has been shown to contribute to glucose intolerance in other mammalian models (7). We discovered that while insulin levels increased in two year old rodents compared to one year old rodents in all cohorts, only the two year old wTERT2 rodents showed an increase that reached statistical significance compared to wTERT (Figure 4D).
Lifespan Improvements Result from wTERT2 Ec- topic Expression in Rodent Models
Not only does the Wookiee species show robust physiology but also long lifespans averaging about 400 standard galactic years (Imperial Census, 3645 ATC). We performed Kaplan-Meier analysis of rodents in all three cohorts to assess whether the wTERT2 gene not only imbues physiological robustness but also longevity. We observed that the lifespan of wTERT2 rodents was significantly greater than that of wTERT or control rodents, leading to an increase in survival of wTERT2 rodents of 24% (P<0.05) compared to control rodents. Rodents with wTERT showed no significant increase in lifespan compared to controls (Figure 4E). We also noted a significant increase in the median survival of the longest lived rodents (90th percentile) in the wTERT2 rodent group compared to the control group, suggesting that the wTERT2 gene may affect maximum longevity. Not surprisingly, the longest lived wTERT2 rodent group surpassed the control group by 13% (P<0.05) in maximum longevity (Figure 4E).
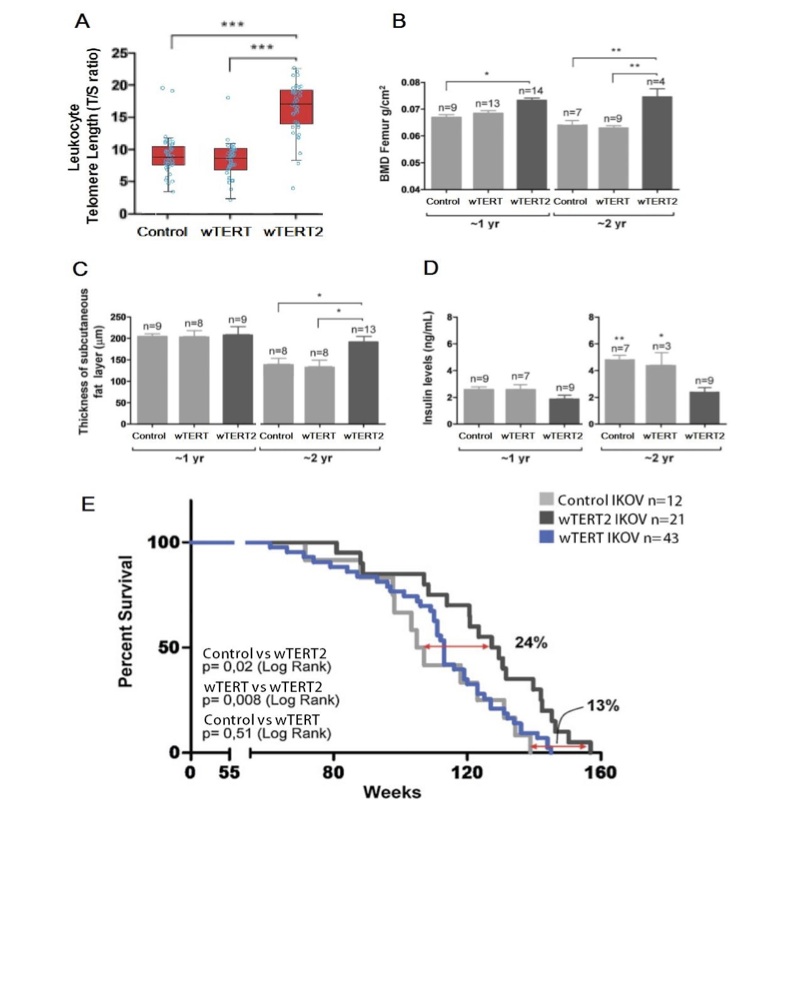
Figure 4 (Click to Enlarge): Delayed aging in wTERT2 transgenic rodents (adapted from de Jesus et al, 2000)
A. Telomere length as measured by T/S ratio, showing significantly increased telomere length in wTERT2 transgenic rodents (p<0.001).
B. Femur bone mineral density (BMD femur) in one and two year old rodents with the indicated vectors. Two-way ANOVA was used for sta- tistical analysis demonstrating that wTERT2 imbues an osteoblastic advantage (p<0.002). Data are given as mean+/-SEM (*p<0.05; **p<0.01).
C. Thickness of the subcutaneous fat layer at the time of death in tissues fixed immediately post-mortem. Two-way ANOVA was used for sta- tistical analysis demonstrating a significant effect of wTERT2 (p<0.04). Data are given as mean+/-SEM (p<0.05).
D. Insulin levels in one and two year old rodents after animals were fasted for at least 8h. Data are given as mean+/-SEM (*p<0.05; **p<0.01).
E. Kaplan-Meier Analysis on transgenic Ikov show longer lifespan in wTERT2 mice compared to wTERT and iTERT (p<0.05)
To address any undesirable effects of our Zinc- Finger Nuclease strategy, we thoroughly examined rodents for several pathologies at time of death. It has been previously reported that there exists a propensity in TERT transgenic animal models for development of malignant adenocarcinomas and increased incidences of benign hyperplasias (7). Neither the wTERT or wTERT2 rodents in this study exhibited increased incidences of cancer compared to control rodents.
No significant difference in telomerase expression was detected between wTERT and wTERT2 rodents using TERT RT-qPCR (Figure 5A). However, via Telomerase Repeat Amplification Protocol (TRAP) assays we were able to show that the wTERT2 rodents displayed significantly more telomerase activity compared to both wTERT1 and iTERT controls (Figure 5B). We suspected that due to wTERT and wTERT2 sharing the same promoter in our transgenic rodents, the expression of the proteins would not differ. Nevertheless, the above TRAP assay derived data suggests that wTERT2 is a more active reverse transcriptase compared to both wTERT and iTERT. This may help to explain why short-lived Wookiees have lifespans similar to that of other Imperial races, roughly 100 std galactic years compared to 400 in normal Wookiees.
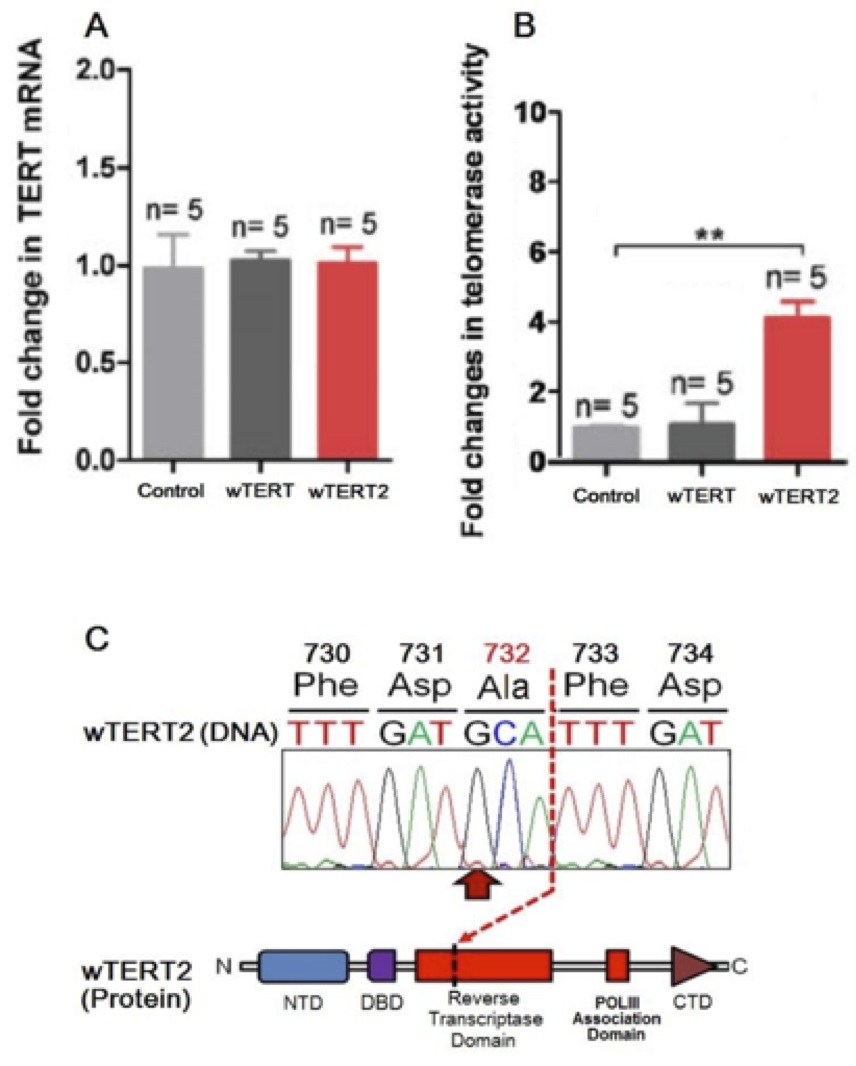
Figure 5 (Click to Enlarge): Expression and catalytic activity of wTERT2 is neces- sary for the elongation of telomeres (from de Jesus et al, 2000)
A. Fold change in TERT mRNA levels in wTERT or wTERT2 rodents compared to controls. Values are nor- malized for acidic ribosomal phosphoprotein P0 (RPLP0) and are represented as the change in mean+/-SEM. Student’s t-test was used for statistical analysis.
B. Telomerase activity (measured through TRAP assay) in peripheral leukocytes in wTERT or wTERT2 rodents compared to controls. Student’s t-test was used for statistical analysis. (** p<0.01).
C. Mutational landscape of the most common wTERT2 non- synonymous substitution as derived from targeted exome sequencing.
Targeted Exome-Sequencing of the wTERT2 Locus Potentially Reveals Causative Mutation in Prematurely-Aged Wookiees
To establish why expression of wTERT2 is almost non-existent in any Wookiee population under the age of 80 but rapidly upregulated after the age of 80 and what the consequences of this might be, we con- ducted targeted exome sequencing on the wTERT2 genomic loci of advanced-aged Wookiees compared to a cohort of age-matched phenotypically normal Wookiees. Exome Data was aligned to the Wookiee wookiee consensus genome, confirmed by Sanger sequencing, and compiled for prematurely-aged Wookiees.
As evidenced by the overwhelming number of prematurely-aged Wookiees in the cohort being categorized with a non-synonymous mutation at base 2196 causing a shift in residue 732 from threonine to alanine or serine of the wTERT2 gene (Figure 5C), homozygous loss-of-function mutations of the gene must have a causal role in the development of premature senescence in the Wookiee species. Though not all prematurely-aged Wookiees presented with the A to G/U substitution, the vast majority showed a substitution, resulting in translation of a non-functional protein product.
Discussion
In this study, we characterize a novel telomerase reverse transcriptase, wTERT2, essential for upkeep of telomere length in Wookiees past the age of maturity at 80 years. The wTERT2 protein, previously known as RTL4 (reverse-transcriptase-like protein 4), is part of the family of reverse-transcriptase-like proteins. Although wTERT2 is expressed in the nucleus, RTL2 and RTL3 are expressed in the midi-chlorians (8), and the RTL family of proteins were previously thought to be involved in the replication of midiDNA (8). The wTERT2 enzyme shares high homology with previously characterized wTERT, especially in the reverse transcriptase and RNA binding domains (8). As such, we expect protein-protein interactions of wTERT to be comparable to those of wTERT2.
Employing transgenic Kashyyyk Ikov rodents as animal models for functional analysis of wTERT2, we observe that replacing wTERT2 in place of iTERT in the Ikov genome confers anti-aging qualities such as lengthened leukocyte telomeres, delay of age-related comorbidities, and extended lifespan. These data suggested to our research group that aberrations in the wTERT2 gene could have a physiological link in the early senescence of Kashyyykan Wookiees.
Returning to our cohort of Wookiees afflicted with IPF, we performed targeted exome sequencing on the wTERT2 locus and observed a common A2196X single nucleotide substitution. This mutation most frequently resulted in the substitution at the 732nd residue of threonine to alanine or serine. In our cohort, 96% of IPF-afflicted Wookiees have one of the above mutations. We had previously found that when the wTERT2A2196X was substituted into Ikov, none of the animals survived past the embryo stage (data not published), with an inner cell mass (ICM) significantly less than that of a wildtype rodent. This is consistent with previous rodent studies in which TERT was knocked out in the embryo, indicating that the wTERT2A2196X mutation results in a loss of function phenotype. Since Ikov do not have a redundant TERT as Wookiees do, the mutation in these animals is an embryonic lethality.
The above data suggest that wTERT2 plays a vital role in the aging process of Wookiees above the age of 80. The strong correlation of wTERT2A2196X with accelerated aging and IPF suggests an association that needs to be further elucidated in a longitudinal study. One area of interest is the phenotype the mutation manifests molecularly. Our data show single copy mutation in the diploid Wookiee genome results in the prematurely-aged phenotype. Because of the low penetrance of the phenotype in the 50 million total Wookiees on Kashyyyk, the implication is that the mutation results in a haploinsufficiency where a single copy of wTERT2 is insufficient for normal phenotype. This study does not however exclude the possibility that the mutations described may present in a dominant negative fashion.
An important direction for future studies is determining the method by which wTERT2 becomes activated in adult Wookiees. The wTERT2 gene may be awoken from dormancy in response to critically short telomeres, or be tightly regulated at the genetic or even epigenetic level until the Wookiee approaches 80 years old. More functional and also need to be performed to assess the reverse transcriptase activity of wTERT2 via chromatin immunoprecipitation (CHIP) and DNA docking assays. Furthermore, structural studies would be beneficial in confirming the structural, similarity between wTERT and wTERT2, which would help elucidate functional differences
Wookiees with IPF live an average of 30 years after diagnosis. On a human scale, this appears to be a slowly progressing disease. However, in Wookiees this represents a reduction of life of more than 70%. There is no cure to this affliction, but this paper provides a novel target in wTERT2 for further functional studies, epigenetic studies, and potential therapeutics. The IHRI is proud of the progress that has been in made in the field of human anti-aging medicine, especially with regards to the success of the lentiviral-based hTERT gene therapy developed on Youst Station. It is possible similar breakthroughs can be achieved by focusing our efforts on wTERT2. The end goal of such research could dramatically affect the productivity for the Empire’s Wookiee labour force.
Materials and Methods
Population
Study participants consisted of age-matched IPF-afflicted and IPF-unafflicted Wookiees between the ages of 19 and 150. The interquartile age range is between 64 and 114. Informed consent was given via proxy by the Imperial Governor of Kashyyyk, Hindane Darcc. This study was approved by the Coruscant Im- perial College Clinical Research Ethics Board, and is in full compliance with IHRI Research Ethics Guidelines. Our cohort was split into four groups based on age cutoff and IPF status. 134 prematurely aging, IPF- suffering Wookiees under the age of 80 were age- matched with 151 Wookiees that showed no signs of accelerated aging. 163 prematurely aging, IPF-afflicted Wookiees over the age of 80 were age-matched with 198 Wookiees that showed no signs of accelerated aging. The Rwook subspecies of Wookiees were the most heavily represented in our cohort. Of the 646 Wookiees in our study, 44 were albinism-related subspecies. Excluding non-Rwook samples yielded no change in the significance of our comparisons. Also, no new significant trends emerged when non-Rwook samples were analyzed as their own group. Clinical information and blood were collected at Rwookrrorro General Hospital and Kashyyyk Royal City Hospital as part of the work- force annual physical.
DNA Microarray
Total RNA was isolated from whole blood. 3mL of peripheral blood was collected through venipuncture into ethylenediaminetetraacetic acid (EDTA) tubes, and frozen as whole blood at -80oC. Total RNA was isolated from 3mL of freshly thawed whole blood using the BloodRNA kit from Alderaan Genomics (Cat#345987). 1000ng aliquots of the RNA samples were reverse transcribed using the One-Cycle cDNA Synthesis Kit (Corellia Technologies, Cat# 6984). The samples were either Cy3- or Cy5-labelled using the EZAmp Fluorescent Amplification Kit (Geonosian LifeCorp, Cat# 243). Cy3- and Cy5-labelled samples were combined and hybridized to the Ikov Oligo Microarrays as per manufacturer instructions (Geonosian LifeCorp, Cat# 4213). The microarrays were quantified post-wash using the PlateScanner from Nubian BioSys- tems (Ref# 56566). Cy3 and Cy5 dye-swapping was performed on a complete set of duplicate samples. PlateScanner software was also used to analyze expression data. Candidate genes that were found to be up-regulated or downregulated in association with one of the cohort groups (p=0.005) were screened using BLAST against the IHRI Kashyyyk Genome Databases (Wookiee wookiee Consensus Genome therein) for aging-related genes
ZFN Rodent Model Development
Zinc-Finger Nucleases (ZFNs) specific to the iTERT locus were used to create transgenic rodents with GFP-tagged wTERT and wTERT2 genes instead of iTERT. Due to sequence homology between wTERT and iTERT, expression of wTERT-GFP and wTERT2-GFP should be similar between the two transgenic mice. PCR fusion was used to construct wTERT-GFP and wTERT2-GFP as previously described (7). ZFN expression plasmids were created in-house using Rwook embryonic kidney cell line hek2α DNA, and validated for activity by transfection into Ikov carcino- ma cell line 4B4. Transfected cells lines were sequenced with Kashyyykan TERT primers FO2555 and FO2556 as previously described (7).
Ikov were housed in static cages in a room with a 14hr/10hr light/dark cycle. Water was provided as needed, and the Ikov were kept on a Western-type diet (Tatooine Rodent Genetics Inc, Cat# 254). Only female Ikov were used, and all mice have >99% (C56BL/6 background). Pregnant Mare Serum (PMS) was injected at 3IU/animal into 6-week old Ikov, followed by Human Chorionic Gonadotropin (hCG) injection after 48 hours at 5IU/animal. Single-cell fertilized eggs were harvested for microinjection 15 hours post-hCG injec- tion. Microinjection of 5ng/ul ZFN mRNA was per- formed, and injected eggs were implanted in pseudo-pregnant Ikov provided by Tatooine Rodent Genetics Inc. Integration and expression of transgenes wTERT- GFP and wTERT2-GFP were confirmed by GFP fluo rescence quantification in shaved back skin of rodents at birth and every 6 months thereafter. Aging markers, age-related comorbidities, and Kaplan-Meier analyses were performed on Ikov of all three cohorts: iTERT (wildtype, n=14), wTERT-GFP (wTERT, n=21), and wTERT2-GFP (wTERT2, n=23).
Bone Density
A Dual Energy X-Ray Absorptiometry (DEXA) scanning instrument (Kaminoan Medical Technologies, Ref# 41) was used to measure bone mineral density (BMD) in anesthetized Ikov.
Subcutaneous Adipose Thickness
Five 3mm thick skin sections were collected from each Ikov, and 10 random measurements were conducted from each section to create an average value per animal. Sections were stained for hematoxylin-eosin (H&E), and length measurements were made using QuikImage software (Kaminoan Medical Technologies, Ref# 252).
Glucose Intolerance
1 year old and 2 year old Ikov were fasted for 5 hours, followed by intraperitoneal (IP) injection of 50% dextrose solution at 5g/kg body weight. Blood insulin levels were quantified as per manufacturer specifica- tions using an Ikov Insulin ELISA Kit (Nubian Biosystems, Ref# 857)
Leukocyte Telomere Length
Peripheral blood was collected from tail vein in 2 year old Ikov. DNA was isolated from whole blood using the BloodDNA Kit (Alderaan Genomics, Cat#1777). Leukocyte telomere length was measured in triplicate using monochrome multiplex polymerase chain reaction (MMqPCR) as previously described (3), using telg, telc, albu, and albd primers. PCR was per- formed on a BritePlate 480 instrument (Utapau Life Sciences, Cat# 5144). Relative quantitation was per- formed using 10 concentrations of serially diluted standard DNA. Amplification efficiency was between 97% and 100% for both amplicons. T/S ratios were generated by dividing a thousand times the telomere copy number by the single copy gene (albumin) copy number. This MMqPCR technique was validated using fluorescence in situ hybdridization and flow cytometry (flow-FISH). A high correlation between flow-FISH and MMqPCR was observed (R2 = 0.91).
Telomerase Activity
Leukocyte telomerase activity was assayed using a variant of the Telomerase Repeat Amplification Protocol (TRAP) assay on cell lysate isolated from whole blood. Leukocytes were isolated, lysed, and assayed using the TaunTAGGG PCR ELISA kit (Hoth BioMedical, Cat# 695) in accordance to manufacturer’s proto- col. Relative telomerase activity percent (RTA%) was determined following instructions from the TaunTAGGG protocol. Messenger RNA levels were also measured of wTERT, wTERT2, and iTERT in transgenic Ikov. Total RNA was isolated from 3mL peripheral blood, and converted into cDNA as above. Primers for wTERT were used as follows from a previous publication: forward 5’- TGACACCTCACCTCACCCAC-3’ and reverse 5’- CACTGTCTTCCGCAAGTTCAC-3’. Ikov acidic ri- bosomal phosphoprotein P0 (RPLP0) was used a housekeeping gene. RPLP0 primers are forward 5’- GGCGACCTGGAAGTCCAACT-3’ and reverse 5’- CCATCAGCACCACAGCCTTC-3’. RPLP0 transcript levels were used to normalize the signal from wTERT, wTERT2, and iTERT.
Targeted Exome Sequencing
DNA was extracted from peripheral-blood leu- kocytes as before, and genomic DNA was subjected to array capture with the PreciseSelect Exon Kit (Bespin Biosystems) according to the manufacturer’s instruc- tions. Adapters were ligated, and 2 × 69 bp paired-end sequencing was performed on an Illumina HiSeq 9000 at the Coruscant Life Sciences Institute. Each sample produced on average 10 Gb of sequence covering 85% of the target sequence with at least 300 reads. Sequencing reads were aligned to the Reference Wookiee ge- nome (wg19, IHRI Genome Browser) with the Vizier-Pellaeon Aligner (v.0.5.7), consensus and variant bases were called with the Genome Analysis Toolkit, and variants were annotated with ANNOVAR.
To validate the A2196G (p.Thr732Ala) mutation, we amplified individual genomic DNA samples with primer pair FO2234 and FO2234 and used Sanger sequencing with FO2823 and FO2824 primers (see Table S1 for oligonucleotide sequences). To validate the A2196U (p.Thr732Ser) mutation, we amplified individual genomic DNA samples with primer pair FO2618 and FO2619 and Sanger sequenced them with FO2221 and FO2223.
References
1. Bernardes de Jesus, B., Vera, E., Schneeberger, K., Tejera, A. M., Ayuso, E., Bosch, F., & Blasco, M. A. (2012). Telomerase gene therapy in adult and old mice delays aging and increases longevity without increasing cancer. EMBO molecular medicine, 4(8), 691-704.
2. Bièche, I., Noguès, C., Paradis, V., Olivi, M., Bedossa, P., Lidereau, R., & Vidaud, M. (2000). Quantitation of hTERT gene expression in sporadic breast tumors with a real-time reverse transcription- polymerase chain reaction assay. Clinical Cancer Research, 6(2), 452-459.
3. Cawthon, R. M. (2009). Telomere length measure- ment by a novel monochrome multiplex quantitative PCR method. Nucleic acids research, 37(3), e21- e21.
4. Gomes, N., Ryder, O. A., Houck, M. L., Charter, S. J., Walker, W., Forsyth, N. R., … & Wright, W. E. (2011). Comparative biology of mammalian telo- meres: hypotheses on ancestral states and the roles of telomeres in longevity determination. Aging cell, 10(5), 761-768.
5. Calrissian, L., Bink, J. J. (3644) On the Phylogeny of Kashyyykan Species. Kashyyykan Phylogenetics, 4(6), 23-24.
6. Cody, C., Fel, J. A Comprehensive Review of Kashyyykan Rodents. Mid Rim Biology, 2(6), 959-1001.
7. Horn, C., Terrik, M. The role of Telomere and Te- lomerase on Cellular Aging in Species of the Galaxy. Imperial Journal of Biology, 6(11), 7733-7744.
8. Jinn, Q. G., Kenobi, O. W. The effect of Reverse- Transcriptase Analogues in Midi-Chlorian Biology. Imperial Science, 8(9), 432-455
