DENATURING GRADIENT GEL ELECTROPHORESIS (DGGE): AN OVERVIEW
Imagine a scenario, where you have a patient with a life-threatening infection. Such a patient might require prompt antimicrobial therapy. However, in clinical microbiology, there is this constant race against the clock in identifying these disease-causing organisms so that an effective treatment can be provided. In fact, often, it is only after the death of a patient that the disease-causing organism is identified. This is primarily because traditional culture methods can be laborious, and in the case of mixed cultures, other analyses are required to identify microorganisms at the species level. Taken together, the entire process can be time consuming, whereby sometimes over a month is required to accurately identify the disease-causing organism (2).
It is therefore important to find ways to solve these problems that are found in extensive culture methods, and PCR-DGGE (Polymerase Chain Reaction Denaturing Gradient Gel Electrophoresis) is one such way. In essence, PCR-DGGE involves amplification of DNA by genus-specific primers that target 16S rDNA sequences and the subsequent differentiation of this DNA on DGGE gel (1-5).
Introduction
DNA is a molecule responsible for preserving genetic information across species and across time. It consists of a meaningful arrangement of chemicals called nucleotides that are symbolized by “A”, “T”, “C” and “G.” These arrangements tell us a story of each organism or individual, in that the code they produce, represent a detailed instruction book for that organism or individual. Any change introduced in this sequence is called a mutation, and whilst mutations occur randomly, many endure as an organism “acclimatizes” to a new environment. Hence, observing and understanding these mutations would undoubtedly improve our understanding of residual and transient organisms in a variety of environments when observation is difficult. For instance, this would be useful for the monitoring of microbial populations, where culture dependant methods fall short.
Denaturing Gradient Gel Electrophoresis (DGGE) is such a technique that attempts to do this. This molecular biology approach is a fingerprinting methodology that has led to revolutionary changes in many of the traditional routines used in assessing microbial populations (1). Therefore, this paper will briefly explain a number of applications that involve the technique and will also examine how DGGE works in general.
Applications of PCR-DGGE
PCR-DGGE is classified as part of the new discipline of molecular microbial ecology (5). Microbial ecology aims at studying interactions among microorganisms and between microorganisms and their environment. This involves long-term study, which includes various and numerous environmental sample analysis (5). However, conventional cloning, hybridisation and culture methods as mentioned above are not always practical for such investigations. Moreover, these techniques do not provide any information on the dynamics of the microbial populations in complex ecosystems and potential effects of environmental changes on such populations (4, 5, 6). Most importantly, these methods require an extended knowledge of the microorganisms to develop adapted probes that target particular individuals among diverse populations (5).
PCR-DGGE has the advantage of not requiring previous knowledge on microbial populations. It is a fingerprinting approach that can generate a pattern of genetic diversity in complex microbial ecosystems such as gastrointestinal tracts, soils, sediments, deep seas, rivers, hot springs and biofilms (4, 5). A major advantage of this method is its potential to visually profile and monitor changes occurring in various microbial communities, that are undergoing different treatments or modifications. It is a rapid and efficient separation technique of same length DNA sequences (amplified by PCR), which may vary as little as a single base pair modification (4, 5, 6).
Furthermore, PCR-DGGE is a flexible method that allows a unique combination of different approaches for a more accurate identification of, for example, functional genes present in particular bacterial populations or specific bacterial species by using hybridization or species-specific probes (2, 4, 5). This methodology can be utilized in diverse subject areas such as clinical and environmental microbiology and food safety.
PCR-DGGE in clinical microbiology
Examples of DGGE applications in clinical microbiology abound. For instance, DGGE allowed the identification of over 65 Mycoplasma species of human and veterinary origins in less than 24 hours (7). Mycoplasmas are fastidious organisms that require many weeks to culture and other serological tests to be identified. They cause various diseases associated with pneumonia, arthritis, conjunctivitis, infertility and abortion (7). This application of PCR-DGGE could potentially allow considerable savings of time, life and treatment costs.
Another important achievement was that DGGE has been established to have a real potential in screening large number of patients for rapid and reliable identification of deleterious changes in both breast cancer genes BRCA1 and BRCA2. (8). Hence PCR-DGGE allowed the detection of numerous mutations and revealed the existence of unclassified variants that were not reported before (8). This method was also able to demonstrate that animals, such as the Nile crocodile, baboon, red panda, wolf and Taiwan beauty snake, could also be infected by Helicobacter species, bacteria suspected to be responsible for stomach ulcers (9).
PCR-DGGE in environmental microbiology and food safety
PCR-DGGE is also a useful tool in studying complex microbial communities such as the gastrointestinal (GI) tract of food producing animals. Food producing animals are animals raised for milk, meat and egg production. These animals can carry in their GI tract disease-causing organisms throughout the production chain to the retail market and from the retail market to the consumer’s dinner table (10). It is therefore very important to elucidate the exact microbial populations of food producing animals’ GI tract. This will allow a better control of the shedding of deadly bacteria strains into manure which is used as fertilizer for produce such as fruits and vegetables.
The recent outbreak of E. coli on spinach in California is a painful recall that even vegetarians are not safe from the damages caused by a degraded health condition of food producing animals. The need for rapid and accurate methods for screening of total microbial populations in complex ecosystems is more evident than ever. PCR-DGGE has proved to be a powerful tool in assessing total gut microbial populations and was also used to detect previously unknown bacteria species in the GI tract of animals (1, 2, 9, 11). Understanding the relationship between the host and the disease-causing organisms will certainly assist us in defining efficient pathogens control measures. This is of paramount importance in food safety and food processing where quality control and assurance programs necessitate proficient methods to discontinue the transmission cycle of life-threatening microbes.
Last but not least, PCR-DGGE was used to examine complex ecosystems such as river, seas, soils and deep-sea hydrothermal vent (4, 5, 12, 13). Understanding complex microbial populations would certainly help us in rapid decision-making with regard to adequate treatment and other major interventions aiming at making the world a better and safer place to live.
How does it work?
First, DNA fragments from a sample containing multiple organisms are amplified using the PCR technique (to learn more about PCR click the following link.) These amplified products generally entail sequences that are well conserved from organism to organism – for example, sequences for the 16S rDNA are a common choice. This collection of fragments is then subjected to the DGGE component of the procedure.
DGGE is a particular type of gel electrophoresis in which a constant heat (about 60ºC) and an increasing concentration of denaturing chemicals are used to force DNA molecules to unwind. A quick glimpse at electrophoresis tells us that this is a separation technique based on the electrical charge, shape and molecular weight of particulates such as DNA, proteins and RNA (3). In DGGE, DNA, which is negatively charged, is attracted by the positive electrode and forced to migrate through the pores of a polyacrylamide gel. Once it reaches the concentration of denaturing reagents at which it unwinds, it is said to have melted. This determines the melting domains (4, 5), which are defined as stretches of base pairs with an identical melting temperature (4). In other words, base pairs formed by nucleotides A (adenine) and T (thymine), and those formed by C (cytosine) and G (guanine) are chemically melted apart. Basically, what happens is that hydrogen bonding between the base pairs is broken by the temperature and the increasing gradient of denaturing chemicals (urea and formamide) (4, 5). Any variation of DNA sequences within these domains will result in different melting temperatures, thus causing different sequences to migrate at different positions in the gel (4). This provides DGGE with the power to distinguish between mutated and wild type sequences without prior knowledge of what these sequences are, justifying why this method is used to detect mutations within closely related organisms (1, 2).
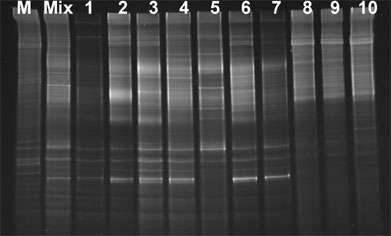
This separation is also aided considerably when a short sequence of G’s and C’s (about 40 nucleotides), often called GC-clamp, is attached to one end of the amplified DNA products (4, 6). This can be done by incorporation of this GC sequence into one of the primers used for amplifying the 16S rDNA fragments.
Conclusion
DGGE is undeniably a valuable approach in screening complex ecosystems on a large scale and in analyzing various environmental samples in a reduced amount of time. Using this technique, diagnosis of emerging infections could become easier and faster, and identification of uncultivable pathogens can also now be facilitated. Although there are limitations, DGGE is an interesting and unique approach that bridges many molecular biology tools together, and its limitations are primarily attributable to the fact that it is still a relatively new technique. If improved,
References:
1. McAuliffe L, Ellis RJ, Lawes JR, Ayling RD, Nicholas RA. 2005. 16S rDNA PCR and denaturing gradient gel electrophoresis; a single generic test for detecting and differentiating Mycoplasma species. J Med Microbiol. 54(Pt 8 ):731-9.
2. Walter J, Tannock GW, Tilsala-Timisjarvi A, Rodtong S, Loach DM, Munro K, Alatossava T. 2000. Detection and identification of gastrointestinal Lactobacillus species by using denaturing gradient gel electrophoresis and species-specific PCR primers. Appl Environ Microbiol. 66(1):297-303.
3. Creighton, Thomas C. 1999. Encyclopedia of Molecular Biology, Volumes 1-4. John Wiley & Sons.
4. Muyzer G, de Waal EC, Uitterlinden AG. 1993. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol. 59(3):695-700.
5. Muyzer Gerard and Smalla Kornelia. 1998. Application of denaturing gradient gel electrophoresis (DGGE) and temperature gradient gel electrophoresis (TGGE) in microbial ecology. Antonie van Leeuwenhoek. 73: 127–141.
6. Sheffield VC, Cox DR, Lerman LS, Myers RM. 1989. Attachment of a 40-base-pair G + C-rich sequence (GC-clamp) to genomic DNA fragments by the polymerase chain reaction results in improved detection of single-base changes. Proc Natl Acad Sci U S A. 86(1):232-6.
7. McAuliffe Laura, Ellis Richard J, Lawes Jo R, Ayling Roger D and Nicholas Robin AJ. 2005. 16S rDNA PCR and denaturing gradient gel electrophoresis; a single generic test for detecting and differentiating Mycoplasma species. J Med Microbiol 54:731-739.
8. van der Hout Annemarie H., van den Ouweland Ans M.W., van der Luijt Rob B., Gille Hans J.P., ¨lle Bodmer Danie, Bru¨ggenwirth Hennie, Mulder Inge M., van der Vlies Pieter, Elfferich Peter, Huisman Maarten T., ten Berge Annelies M., Kromosoeto Joan, Jansen Rumo P.M., van Zon Patrick H.A., Vriesman Thyrsa, Arts Neeltje, Boutmy-de Lange Majella, Oosterwijk Jan C., Meijers-Heijboer Hanne,. Ausems Margreet G.E.M, Hoogerbrugge Nicoline, Verhoef Senno, Halley Dicky J.J., Vos Yvonne J., Hogervorst Frans, Ligtenberg Marjolijn, and Hofstra Robert M.W.. 2006. A DGGE System for Comprehensive Mutation Screening of BRCA1 and BRCA2: Application in a Dutch Cancer Clinic Setting. Human Mutation. 27(7):654-666.
9. Al-Soud Waleed Abu, Bennedsen Mads, W Stephen L. On, Ouis Ibn-Sina, Vandamme Peter, Nilsson Hans-Olof, Ljungh Åsa and Wadström Torkel. Assessment of PCR-DGGE for the identification of diverse Helicobacter species, and application to faecal samples from zoo animals to determine Helicobacter prevalence. J Med Microbiol 52 (2003), 765-771
10. National Research Council. Committee on Drug Use in Food Animals. Panel on Animal Health, Food Safety, and Public Health. Board on Agriculture. Food and Nutrition Board. The use of drugs in Food animals. Benefits and risks. Washington, D.C: National Academy Press; 1999.
11. Gong Jianhua, Forster Robert J., Yu Hai,. Chambers James R, Sabour Parviz M., Wheatcroft Roger and Chen Shu. 2002. Diversity and phylogenetic analysis of bacteria in the mucosa of chicken ceca and comparison with bacteria in the cecal lumen. FEMS Microbiology Letters. 208(1):1-7.
12. Nakatsu Cindy H., Torsvik Vigdis and Øvreås Lise. Soil Community analysis using DGGE of 16S rDNA Polymerase Chain Reaction products. Soil Science Society of America Journal 64:1382-1388.
13. Muyzer G, Teske A, Wirsen CO, Jannasch HW. 1995. Phylogenetic relationships of Thiomicrospira species and their identification in deep-sea hydrothermal vent samples by denaturing gradient gel electrophoresis of 16S rDNA fragments. Arch Microbiol. 164(3):165-72.