BREAKFAST OF CHAMPIONS DOES REPLICATION
To begin with, we’ll start with a chicken scratch drawing of a DNA molecule, which you know is double stranded. My poor pathetic attempt at illustration is therefore going to look like this:

You also know that each strand of DNA is composed of building blocks called nucleotides, and that these nucleotides are always interacting in a complementary manner. For example, A’s are always with T’s, C’s are always with G’s, Beavis is always with Butthead, etc etc etc. Let’s draw them in like so:
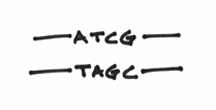
What you haven’t been told at this point is that chemically speaking, the two strands are going in opposite directions. The correct term for this is actually known as anti-parallelism. To denote this, I’ll draw some arrowheads on the DNA strands:
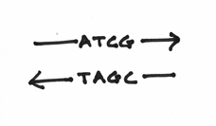
Although, this may seem a little confusing at first, try to picture two lines of square dancers facing each other. In this circumstance, you notice that when focusing on the left or right hands of the row of dancers, the two lines are going in opposite directions. This picture should help:

Your DNA strands are doing something very similar in a chemical sense. The difference, of course, is that instead of dancers, you have your choice of four nucleotides. Furthermore, like the situation of left hands versus right hands, the ends of the DNA strands are also different. One end is known as the 3’ (pronounced 3 prime) end and the other is known as the 5’ end. To the layman, these rather stoic terms are an unfortunate consequence of chemical labeling. So now, our picture should look like this:
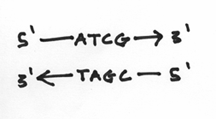
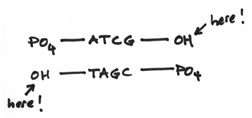
I should reemphasis that the 3’ and 5’ ends are very different from each other. To be more specific, we say that they are chemically distinct from each other. They are as different from each other as apples and oranges. In fact the 3’ end is composed of a hydroxide group and the 5’ end is composed of something known as a phosphate group. These groups look a little like this:
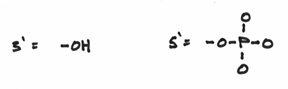
Hopefully, it’s easy to see that they are indeed distinct from each other —even more so than apples and oranges. The hydroxide group being comparatively small and meek, whereas the phosphate group is prominent, overbearing even. This turns out to be a crucial factor because replication is carried out by the activities of a variety of different enzymes which all function by focusing on one DNA end or another or both.
So now, the picture looks like this:
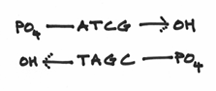
It should also be pointed out that DNA is not really like this flat goofy looking cartoon. As mentioned in a previous chapter, the two DNA strands are actually intertwined around each other in a rather pretty helical fashion. This is where the two strands are wound around each other, sort of like two elastic strings twisted and coiled together. Sort of like this:
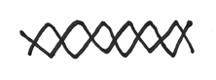
Now that the stage is set, it’s time to introduce the proteins or the enzymes, which are responsible for the actual process of replication. Enzyme is just a fancy word for a protein that is able to facilitate a chemical process. What I’ll do here is to focus on terminology associated with a simple organism like the bacteria, e. coli. However, all organisms, even those as complicated as humans, do more or less the same thing when it comes to doubling their DNA — the principle difference being that unfortunately, the enzymes have difference names and labels.
That aside, the first enzyme for replication in e. coli that we should introduce is, of course, the most important enzyme in the entire process. In e. coli, this enzyme is called DNA polymerase III (or DNA pol III for short), and is essentially the one that is responsible for the actual business of making more DNA. If this entire exercise was analogous to a movie, then this enzyme is the marquee player. It is the Tom Cruise, the Julia Roberts, the proverbial bread and butter of replication. It is, quite simply, the star of the entire process. Instead of drawing a picture of Tom Cruise or a picture of Julia Roberts, I think a picture like this should suffice:

Problem is, if we were to draw this enzyme to scale with a helical DNA molecule (like this),

you’ll notice that the DNA pol III is actually too big to get inside the DNA strands. It can’t go about its business of copying the DNA, because the strands are all coiled up in the helical structure. In other words, there is a serious issue of accessibility. Even our star enzyme, despite its importance, can’t do its job without access to the molecules of DNA it wants to copy. Consequently, the enzyme that inevitably has to act first is one that is responsible for opening up the DNA strand. This enzyme is known as a helicase, and its role is to essentially unwind the DNA molecule, which would look like this:
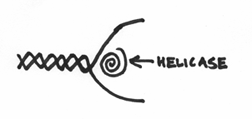
The net effect being the production of a “bubble” of opening where the two DNA strands are pried apart and are subsequently accessible to the whims of the replication machinary.
Curiously, the DNA pol III, which after the unwinding event, can now interact with the DNA molecules, does so whilst attached to a bunch of other enzymes. This attachment is a little like a bunch of buddies hanging out together. The complex actually looks a little like this:

You’ll notice it has the following… (i) two DNA polymerase III’s: which kind of makes sense given the fact that there are two strands of DNA that need to be copied; (ii) one helicase molecule: which also sort of makes sense, because as this replication complex is doing its thing along the DNA molecule, wouldn’t it be handy to have the built-in ability of opening up the DNA molecule as it moves along; and (iii) one new enzyme which is known in e. coli as the primase. However, the purpose of the primase molecule is a little complicated and so to fully comprehend the role of this enzyme, we need to switch gears a little and tell you a bit more about the DNA pol III molecule.
What actually needs to be done, is for us to go over a few mechanisms that all DNA polymerases seem to use. In fact, it’s apparent that every DNA polymerase that has been discovered on this planet:
In fact they all (without exception) seem to follow a two basic rules.
Rule number one states that all DNA polymerases function by adding nucleotides to the 3’ end of the DNA strand. What this means exactly is that a DNA strand can be extended by the addition of new A’s, T’s, C’s or G’s. However, the new nucleotides can only be added to one particular end, namely the 3’ hydroxide group. This is a molecular restraint in that the DNA polymerase can only join nucleotides via this smallish chemical group. This rule can be drawn out like this:
Rule number two states that all DNA polymerases require a primer to function properly. This is probably the most challenging concept that needs to be addressed. If you get through this, then you consider yourself home free.
To simplify the notion of a primer, let’s look at a single strand of DNA, complete with its 5’ and 3’ ends. It should look a bit like this:

Now according to rule number one, a DNA polymerase can extend this single strand chain but only by adding nucleotides to the 3’ end. In effect, you can argue that all of the relevant chemical groups are present for making more DNA. However, the problem lies in the fact that under these circumstances, the DNA polymerase doesn’t actually know what to add. How does it know, whether to add an A, a T, a C or a G? It can’t exactly be a random event, because replication is all about making sure cells receives an identical copy of the DNA code.
Take the following picture:

Under this layout, it should be clear that now, the DNA polymerase has the required 3’ group, AND it also has a template to read and ascertain what those nucleotides should be. For instance, if the nucleotide in the opposite strand is a G, then the DNA polymerase knows it should add a C. If the nucleotide in the opposite strand is a T, then the DNA polymerase knows it should add a A. Hopefully, at this point, you’ll at least agree with the following statement. A DNA polymerase can not do anything with a single strand of DNA. True, it has the right chemistry, but in effect, it does not have the template or instructions needed to define how the chain is extended.
If we redraw the picture. Say like this:

What you’ll notice are two strands of DNA, one long and one short. You’ll also notice that the strands are anti-parallel as discussed earlier. If you focus on the arrowhead, you’ll find yourself focusing on a perfectly situated 3’ group. Here is the end of a DNA strand that is chemically ready to have nucleotides attached. Furthermore, it is also a 3’ end that is located where a template is present on the opposite strand. In other words, everything is in place. The right chemistry, and a means for instructing which nucleotides to add. Again, taken at the simplest level, we can conclude that in order for a DNA polymerase to do its thing, it needs an area of double strandedness.
So,.. the small sequence of nucleotides that has been circled here…
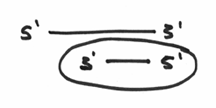
… which makes an area of double strandedness is technically known as a primer. With this all sorted out, hopefully the rule about requiring this primer makes a little more sense, and you can probably guess that the enzyme called the primase may have something to do with this nuance.
Which turns out to be exactly what this primase enzyme is all about. In a nutshell, it is an enzyme capable of making a short sequence of nucleic acids which functions as a primer. A key point that needs to be emphasized, however, is that this primer is made up of RNA, which if you recall, is a molecule that is very similar to DNA in that it is also composed of the representative four nucleotide code. This is actually due to a biological technicality whereby it is possible to make a complementary strand of RNA without the use of a primer (hmmm, think about this for a second). Taken together, the function of the primase should end up looking like this:
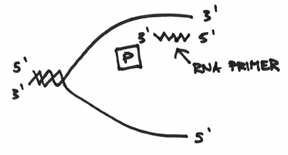
If you’ve been following along, then hopefully you can see that replication from this RNA primer can proceed in a manner that can be drawn like this:
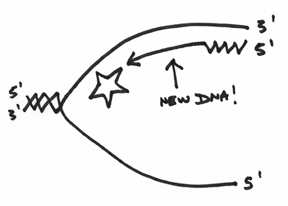
However, it’s wise to pause here for a second, because you have to understand that whilst this top strand is being replicated, the lower strand is also being worked on simultaneously (There are two DNA polymerase III’s attached together afterall). The lower strand is actually a bit messier for reasons that will become clearer as we proceed in this discussion.
Basically, the primase enzyme will also go about preparing a primer for the lower strands. However, if we draw this primer and label the ends in the anti-parallel manner, you can hopefully see a logistical problem in this set-up. Take a look at the following picture, and see if you can find the problem (remember, the DNA polymerases, the helicase and the primase all move as a single unit in one direction, and remember that all DNA polymerases must add to the 3’ end):
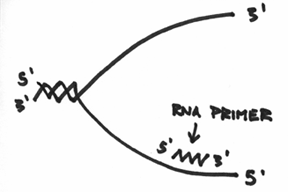
Do you see the problem? Do you see a problem with the direction of the primer? Do you see that the 3’ end of the lower primer is facing the wrong direction?
This is obviously a problem, and it turns out that in order to overcome it, the DNA polymerase will still add nucleotides to the 3’ end, but can only do so for a short distance. To keep it simple, think of it as being able to replicate as far as the enzyme is big, which should look a little like this:
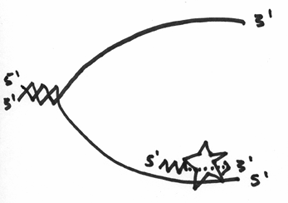
Unfortunately, this doesn’t inherently solve the direction problem, so what ends up happening, is that with this lower strand, the primase has to continually make a primer, and the DNA polymerase III has to continually replicate a little bit. In the end, it should look like this:
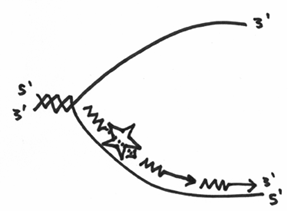
The difference in how each strand gets copied is reflected in why some people call them the leading and lagging strands of replication. One strand is obviously fairly straight forward whereas the other is quite labour intensive.
Anyhow, after this is all said and done, hopefully, you’ll agree with the following statement. That is, we have finally doubled or copied our genetic sequence. However, it should also be clear that the whole thing is a bit messy. For instance, there are bits of RNA everywhere, and the lagging strand is composed of pieces. To address these problems, we have to introduce a few more enzymes.
The first of which is DNA polymerase I, which I will draw as a fish with sharp teeth. This enzyme is special in that, in a nutshell, it is responsible for dealing with the RNA. In a nutshell, its job is to somehow replace it with DNA. In a nutshell, I’ll draw it like this:

DNA polymerase actually has two distinct functions. Firstly, as its name implies, it is a DNA polymerase, meaning that it is capable of extending the DNA chain, but in doing so must follow the same two rules that govern these enzymes. In other words, it must add nucleotides to the 3’ end and it must use a primer as a springboard. Ironically, it is a shitty DNA polymerase. Whereas DNA polymerase III can replicate for several hundred nucleotides, DNA polymerase I has difficulty getting past a few dozen.
Secondly, DNA polymerase I is also an exonuclease. This means it’s capable of degrading or chewing up nucleotides. Which is another reason why I drew a fish with teeth. And not only does it chew stuff up, it does so in a fairly specific manner. To begin with, it likes to start at areas, which are termed as nicks in the DNA. In our picture, this is where the nicks would be:
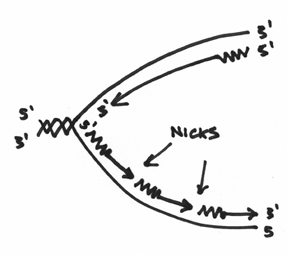
Furthermore, this exonuclease is picky in that it always chews from the 5’ end. Basically it is gunning for that big phosphate group. So that you don’t forget this, I’ve drawn this picture to help you visualize this:
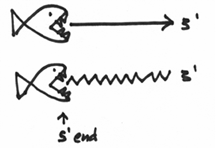
Now, if you take all of this into consideration, you come up with the following mechanism. DNA polymerase I will come in on our replication picture, and zone in on a nick in the strands. Once there, it will begin chewing on the 5’ end, which should look a bit like this:

Don’t forget that this enzyme is also a DNA polymerase, and if you look at the other side of the nick, you will hopefully realize that there is this beautiful 3’ end ready for action. This beautiful 3’ end is right here:
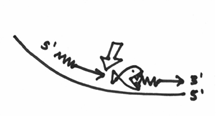
Let’s say that the fish’s ass happens to contain the DNA polymerase function. What therefore happens is that DNA polymerase I will start replicating from that 3’ end, which incidentally fills up the gap that was created by the exonuclease activity. This should nicely demonstrate how DNA pol I achieves its function of replacing the RNA with DNA. This whole step should kind of look like this:

Hopefully, this puts the shittiness of this DNA pol I in perspective. It’s quite biologically pretty because, I hope you can appreciate that DNA pol I doesn’t need to be very good. It’s only responsible for replicating the small region encompassed by that RNA primer.
So,.. after this enzyme has done its thing, you should now agree with the following statement — that you have now doubled your DNA. Of course, it’s still a bit untidy because the strands (especially the lagging strand) are still in bits and pieces. Enter the next and final enzyme, which is called the ligase. This enzyme has only one job and that is to seal all of the bits and pieces together. It fairly analogous to a glue job and essentially your picture will go from something like this:
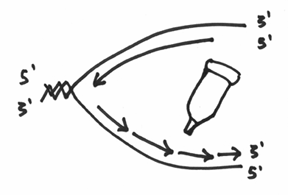
To something like this:
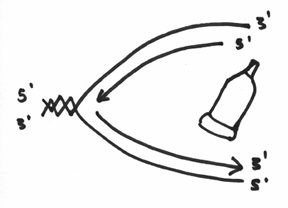
And (drum roll please) VIOLA! You have doubled your DNA. You have made two copies of the same genetic code – which during the process of cell division, will enable each of the two new cells to receive a copy of the genome.
One of the nuances that should be mentioned is that if you examine the entire process, you will notice that each of the DNA sequences is derived from one old strand and one newly synthesized strand. Because of this, replication is often termed semi-conservative, whereby each of the original two strands is read individually to synthesize a new and complementary strand.
Actually, I lied. It’s not quite over. Before, I finally put this whole replication thing to rest, I think it’s also worth talking about one other enzyme, or a family of enzymes, known to scientists as topoisomerases. I like mentioning these enzymes, because I think they do a wonderful job of illustrating just how complicated and elegant nature is, when confronted with a specific job.
What we’ll need to do here is undergo a visual exercise. Let’s say I tell you to hold two fingers up like this:

And let’s say that I have an elastic band. With this elastic band, I will twist and coil it and then place it around both of your fingers. Essentially, this will represent the double helix and will look a bit like this:

If you recall, the first thing that had to happen was for a helicase enzyme to come in and open up that helix structure. Let’s say that I am the helicase, and I come in and grab hold of the two strands of your elastic band and pry them open. It should make a little bubble and should look a little like this:
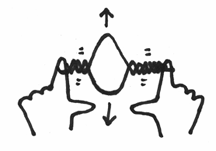
Can you see that under these circumstances, the helix on either side of the opening will be actually twisted even more. It would be like taking your replication fork, grabbing hold of each strand, and like the helicase forcing an opening like this:
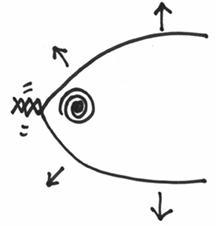
Do you see that this will cause a further tightening of the coil along the helix?
This is actually very bad for the DNA molecule, as this twisting can cause a lot of structural stress. So much so, that the DNA molecule is in very real danger of snapping – which you can imagine would be a very bad thing to happen during replication.
Topoisomerases are enzymes that are designed to take care of this problem. These enzymes can actually detect these areas of high structural stress, and zone in on them. Not only that, but whilst they are at these areas, they will then cut both strands in the DNA complex. Remarkably, they will then hold on to all four ends of the cut, and in a very controlled fashion, unwind to alleviate the stress. Finally, they will also behave like ligases and stick back the correct ends together again.
This is nothing short of amazing, and hopefully you can see that these enzymes play an important role. As the DNA is opening up for replication, there will always be an issue of structural stress, which is always addressed by the actions of these remarkable enzymes.
Anyway… taken together, that, Mr. Trout is what replication is all about.