PRELIMINARY PHYLOGENY OF MAGICAL AND ORDINARY CREATURES: EVIDENCE OF A RECENT DIVERSIFICATION
Annals of Praetachoral Mechanics (2016). Vol 2. Advanced online publication. download pdf
ABSTRACT
We use phloygenetics to accurately describe the relationships between various taxa and elucidate patterns in evolution that leads to the great diversity of life we experience today. The events of “first contact” with the wizarding world has led to a stunning variety of eukaryotic life previously hidden from human contact now available for study. This paper seeks to sample, sequence and assign 46 of these new species into the phylogenetic tree utilizing next-generation methods. The resulting tree incorporates these new taxa with a strong morphological correlation between similar, previously cataloged species. There are a few exceptions in the pattern of phylogenies, namely with the inclusion of a new monophyletic group with a high rate of speciation within the Gibbon clade as well as evidence indicating that the Dodo bird might still be extant, reclassified as Diricawl by wizards.
Keywords: Magic, Creatures, Zoology, Magizoology, Phylogenetics
Introduction
Phylogenetics is a powerful tool for visualizing the relationships between various clades that make up all organismal life. While perspectives regarding the fundamental selection process dictating a clade’s position on the tree have shifted, from morphometrics to genetics [1]; there is no doubt that the eventual goal of this field is to understand an organism’s ancestral pathway and determine the degree of relatedness of one organism to another. As part of the discovery and cataloguing process of a new species, a thorough phylogenetic analysis must be performed in order to determine where on the tree the species is to be placed.
The events of “First-Contact” in which the wizarding world exposed itself to general humanity, and the corresponding declassification of wizarding activity, has resulted in a massive influx of new potential for scientific inquiry. Amongst these revelations is the host of new magical species, organisms that exhibit unnatural behaviour that were sequestered from human eyes by various means [2]. These new species have arisen throughout the globe and marks a surge of possible new discoveries by taxonomists.
While these organisms all exhibit extraordinary magical properties, they are still fundamentally organismal, exhibiting morphological characteristics similar to known taxa [3]. It is possible to obtain blood and tissue samples with the aid of wizards [4]. There has been success in deriving gene sequences from these samples [5] and while there are a multitude of genes not expressed in any known organism, the remaining genes do have real-world counterparts [6]. This similarity invites classification and cataloguing via phylogenetics. Wizards have no need for scientific names (a rough method for classification exists [7]) and consequentially no phylogenetic tree exists for these organisms.
In joint collaboration with the Muggle Liason Office and the Department for Regulation and Control of Magical Creatures of the Ministry of Magic, our goal is to sequence all available biological samples of magical creatures using next-generation sequencing and to determine the orders and families, if any, these species belong to. The samples are gathered from the personal collection of the magizoologist Newt Scamander. We then conduct a phylogenetic analysis on the genes and construct an updated phylogenetic tree with these inclusions.
Materials and Methods
Taxon & Sequence Sampling
Magical taxa were chosen based on those samples contained in Newt Scamander’s personal collection, as well as those found in the Magizoological Museum at the Ministry of Magic. Some samples could not be used, as they were too degraded for DNA extraction. Non-magical taxa were chosen based on descriptions given in Fantastic Beasts & Where to Find Them [9]. The animal described in the book as being most related to a magical-animal was sampled. In addition, some representative taxa were chosen as potential outgroups. For a full list of taxa sampled, see Table 1.
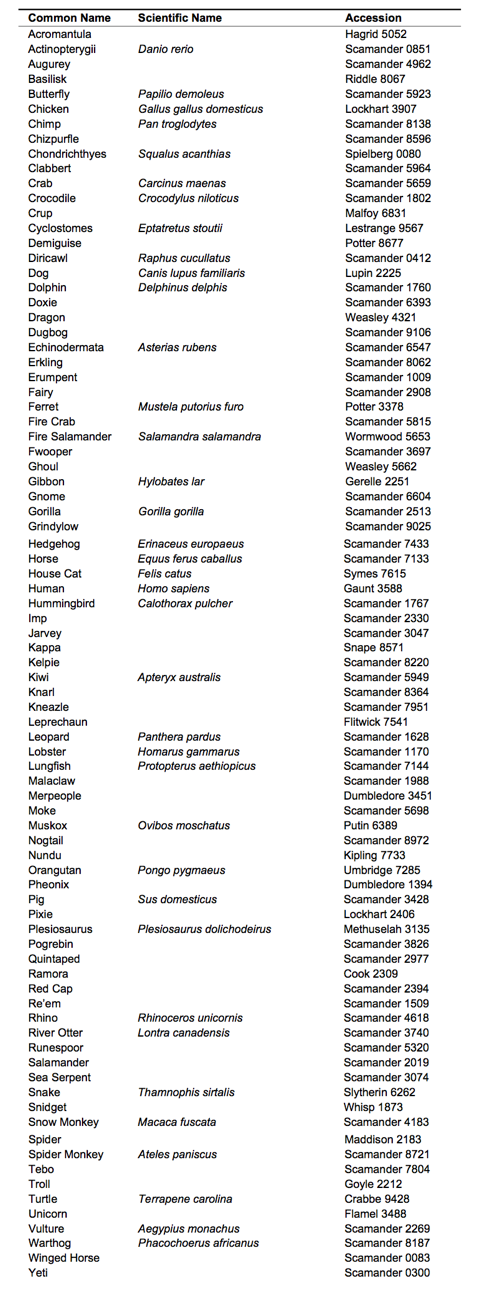
Table 1. (Click to Enlarge) List of taxa with accessions used in all analyses. Accessions are stored in the Magizoological Museum, Department for Regulation and Control of Magical Creatures at the Ministry of Magic. Magical creatures have not yet been assigned a scientific name. Whole clades such as Actinopterygii were used in the tree for simplicity but are listed here with the species used under Scientific Name for analyses.
Genes were chosen based on their commonality among all Eukaryotes. This was done because we weren’t sure of any of the phylogenetic placements of these species and their corresponding genetic makeup. A clade-specific gene may have not existed in the taxon sampled if it wasn’t a member of that clade. Therefore, in the interest of inclusivity, Eukarya-wide genes were chosen. These genes include those involved in the cytoskeleton and DNA replication, as well as others. For a full list of genes used, see Table 2.
Table 2 (Click to Enlarge). List of genes used in all analyses. Genes were chosen based on ubiquity within Eukarya. Accessions are stored under each species name in the database of the Department Maginomics at the Ministry of Magic.
DNA Extraction & Sequencing
DNA was extracted from samples using a DNeasy kit (from Qiagen). Samples were typically taken from muscle or other macroscopic tissue available in Scamander’s collection. Extracted DNA was then sent to the new Ministry of Magic department – The Department of Maginomics. This department is much like any other biology department in a muggle university except with slightly more goblins. Library preparation was performed on extracted DNA and then sequenced on an Illumina HiSeq2000 using a whole genome shotgun approach with paired-end reads.
Raw sequence reads were assembled using CLC Genomics Workbench v.5 (CLC Bio) using a de novo assembly algorithm. Contigs produced from this process were not assembled. Instead, genes of interest were retrieved using tBLASTx. These retrieved genes were placed in an alignment for phylogenetic analysis. Contigs will be assembled in a future paper concerning genomes of magical animals.
Phylogenetic Analyses
Genes were aligned using ClustalW. The resulting output was manually aligned using BioEdit and Mesquite to remove and repair non-homologous alignments. The aligned matrix was partitioned using PartitionFinder into a gene by codon partition. This partitioning scheme was the optimum partitioning scheme as determined by jModelTest. Trees were assembled using a maximum likelihood (ML) approach implemented in RAxML v 8.0.0. The specific settings used were ML + Slow Bootstrap at 100 runs with 500 repetitions. The best tree as determined by RAxML was visualized using FigTree.
Results
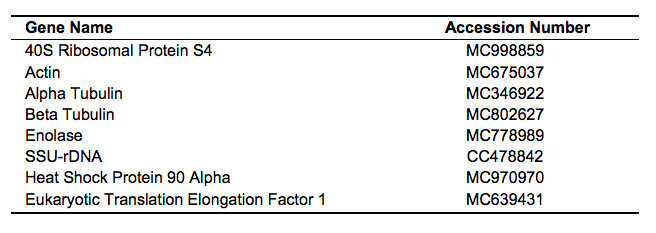
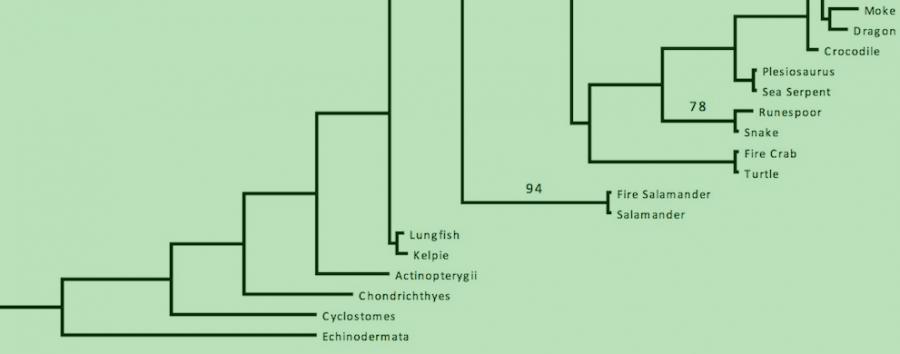
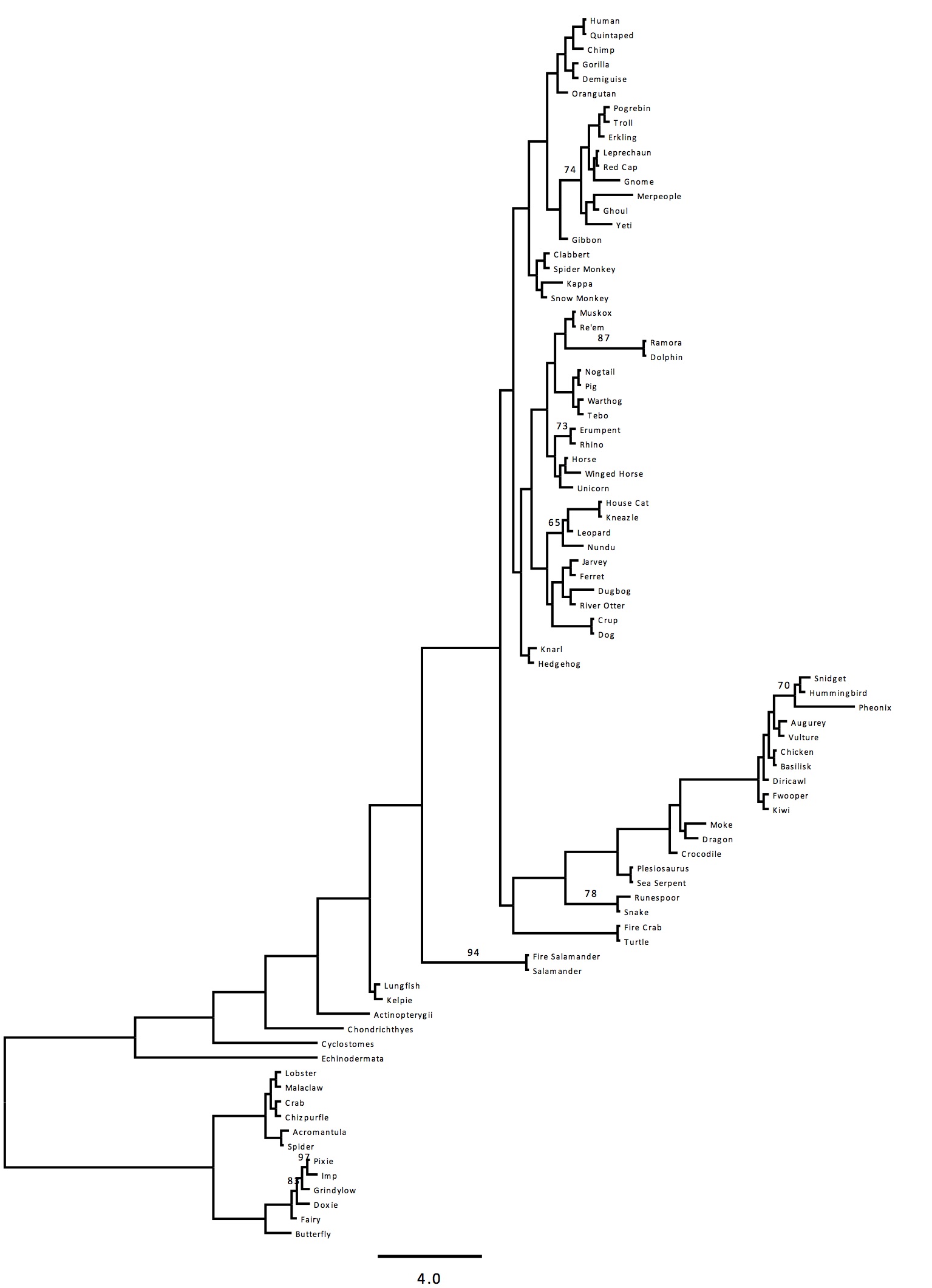
Here we report the first combined phylogeny of magical and ordinary organisms using an eight gene set (Figure 1). All magical creatures sampled appear with Animalia, specifically Vertebrata and Arthropoda. This isn’t to say that all magical creatures are necessarily animals, just the ones sampled in this study. Most magical animals sampled were closely related to another ordinary animal. This is consistent with descriptions given in Fantastic Beasts & Where to Find Them (CITE). There were, however, exceptions to this. For example, two clades of comprised solely of magical creatures were discovered. This includes the Gibbon clade and the extension to the Butterfly clade. Bootstrap support is strong, with most clades receiving 100% bootstrap support. There is moderate support for the Gibbon and Erumpent-Rhino clades. There is poor support for the Feline clade (Nundu, Leopard, Kneazle, House Cat).
Branch lengths are typically short, due to the apparent close relation of each of these animals. There are exceptions to this in the Merpeople, Kappa, Dugbog, and Pheonix. While some rate increases would be expected from a sudden shift to a magical lifestyles these clades show even higher rates.
Figure 1 (Click to Enlarge). Phylogeny of magical and ordinary animals. Taxon names are common names due to the lack of a scientific name for magical animals and for ease of access. For a complete list of scientific names, see Table 2. The tree was inferred using a maximum likelihood (ML) method in RAxML on a partitioned dataset. Bootstrap support values are indicated beside branches; unlabeled branches indicate a support value of 100%. Scale bar indicates substitutions per site.
Discussion
Recent Diversification
Most of these magical creatures share morphological characteristics with many modern taxa, and have branched only recently, and in many cases exist in the same clade with only their sister taxa. While much more study is required to determine other distinguishing differences in the genetic sequences, it is possible to combine all divergent traits under the umbrella of magic, as they express themselves phenotypically as such[8]. Given this assumption, and the relatively equidistant levels of genetic change expressed by all the taxa assayed, there appears to be evidence of a single event in time exposing all the present day taxa to magic, and thus creating the branching events shown in Figure 1. This merits further study, and would be refuted by the presence of magical taxa that express magic at an earlier stage of genetic divergence than ours.
The Dodo Bird
Our analysis and genetic sequencing of the magic flightless bird Diricawl yielded a genetic match with the presumably extinct species Raphus cucullatus or Dodo bird. A BLAST search of the Diricawl genome returned a match catalogued for the Dodo. This return indicates that the Diricawl has a higher correlation coefficient with the dodo than any other of our analyzed taxa. This merits further study, but it is the opinion of the researchers that the existence of the Diricawl refutes the assertion that the Dodo bird is extinct.
Gibbon Clade
Analysis of nine of the magical taxa in the Gibbon clade via ClustalW found sequences unrelated to most known species. Indeed, matches for these particular regions were only present amongst the magical taxa themselves, establishing this group as a new clade. But as we were using an abundantly widespread taxon-rich analyses, we discovered that this new clade is part of a monophyletic group that includes the gibbon and their common ancestor. This group is distinct amongst the rest of the findings as they exhibit wide morphological diversity, as well as the most prolific branching events within magical species that we have seen. It appears that while the rest of the magical species (such as the Gorilla and Demiguise) only exhibit a single branching event before reaching present day, this group (Henceforth dubbed Bogeyreta), has undergone multiple morphological deviations post-magic introduction. It is unclear what circumstances caused this relatively rapid speciation and migrations of these taxa (The Yeti sample, we are told, comes from the mountains of Nepal, while the Leprechaun seems to exist primarily in Ireland). But Bogeyreta is the most taxon-rich monophyletic group encountered so far, and further study is warranted to explore the circumstances behind such rapid evolution.
Problems
As our tree does not measure time intervals, only genetic differences, we cannot describe with certainty at what point in time the magical taxa diverged from their modern counterparts. Also, while our tree was constructed using genetic determination, held by many to be more accurate than the morphological counterpart, the relative correlation between morphological characteristics, the obfuscating factor of magical genomic sequences yet to be properly classified and the relative abundance of separate magical branching events in different species all potentially mask the accurate placement of the taxa on the phylogenetic tree. However, we have taken steps to reduce these inaccuracies by utilizing the most high-throughput techniques available in order to assay sequences with a great degree of accuracy, and are confident that our phylogeny is robust.
Conclusions
The next goal for our team is to organize an excursion into areas of the globe previously inaccessible via magical intervention and conduct a more thorough study on the various taxa present therein. It will be interesting moving forward to see the progression of the Chordate section of the phylogenetic tree, a section that sees rare growth in modern times. In addition, completing genome sequences of these taxa could give us more insight into the proposed singular magical event.
Acknowledgements
The authors would like to thank the Minister for Magic, Kinglsey Shacklebolt, for allowing us into the various magical departments for our experimentation. In addition the authors would like to recognize the help of our molecular biology mentor, Drinking Buddy, as well as Skittles.
References
1. Rédei, George P (2008). Phylogenetic Tree. Encyclopedia of Genetics, Genomics, Proteomics, and Informatics, 1491.
2. Canada. Department of Wizarding Relations. Procedures to prevent Muggle exposure to magical creatures. By R.G Dolamine and G.T Hunz. Ref # 5561-786 January 15, 2012.
3. Geoffery, Randall J. et.al (2014). Similarities of morphological characteristics between Pan paniscus and Wizardensis kappa. Zootaxa, 4110(34), 1360-1362.
4. Reizemenn, K., Humperbuckle, R., & Phillibus, J. (2013). Sample acquisition methods of living Wizardensis demiguise tissue with minimal magic contamination. Nature Methods, 13(2), 122-124.
5. Reizemenn, K., & Jonas, B.(2013). Pyrosequencing of Wizardensis demiguise β-Galactase gene. Nature, 3455(12), 1426-1429;
6. Ionis, T. Y., & Rudd, P. (2013). Different But Same: A study of analogous gene patterns in magical and non-magical organisms. Journal of Magical Biology, 1(4), 312- 315.
7. Luky, T. H., et al. (2012). Interim Classification methodology for magical creatures. Nature, 3401(20), 123-137.
8. Grey, D. et al.(2013). Classification of magic phenotype for future research application. Nature 1232(11), 12- 257.
9. Scamander, Newt. Fantastic Beasts & Where to Find Them. England: Obscurus Books, 1927. Print
Appendix
Genetic sequences for all genes will be provided upon request. Due to magical laws, we are unable to publish them on GenBank.
Editor’s notes: This is the first of many Harry Potter themed papers that will be published over the year. Note that you can submit pieces to the SCQ, as these will be published along side Volume 2 of the Annals of Praetachoral Mechanics. Details on the journal can be found here. Submissions instructions for the SCQ can be found here.