HOW TO MAKE A HAIRLESS WOOKIEE: IDENTIFICATION AND FUNCTION OF DE NOVO GLABR GENE IN WOOKIEE WOOKIEE
(This is the THIRD paper on a special issue on Wookiee science. You can read the first here, and the second here).
Annals of Praetachoral Mechanics. (2014). Vol 1. pp29-38 pdf download.
ABSTRACT
We present evidence for the de novo origin of the Wookiee wookiee protein-coding gene GLABR since their divergence from humans. This gene has no protein-coding homolog in any other genome but its presence is supported by evidence from expression and hybridization data. Furthermore, other near-human species such as Zeltron, Chiss, and Sullustan share the human ortholog of this locus, which supports the inference that the ancestral sequence was noncoding, and that the GLABR has de novo origins in the Wookiee. This GLABR gene was further characterized by a conditional knockout experiment as well as an in situ hybridization on sectioned epithelial cells in order to better determine its function. Our results show that the GLABR gene is responsible for robust hair growth in Wookiees, and that inactivation of this gene results in reduced androgenic hair growth or hairless phenotype.
Keywords: GLABR, hair gene, hair loss, Wookiee wookiee
INTRODUCTION
New Order Laboratories is a subsidiary of the former X1’s Fortress. Originally established as a cloning laboratory, it has since expanded to include an active research and development program. At New Order Laboratories, we aim to expand on the original Wookiee wookiee cloning program, and produce innovative wookiee research to aid the Dark side in their endeavor to rule the galaxy.
Wookiees are favorable to the Dark side due to their destructive bouts of sheer rage, and physical strength. As a whole, wookiees possess several other traits conducive to warfare beginning with the fact that they are a tree dwelling species, an attribute denoted by their species name Wookiee, which translates to “People of the Trees.” As tree-dwellers, Wookiees evolved to have skillful hands and claws. Wookiees also possess fangs and a highly advanced sense of smell.
A distinguishing feature of Wookiee wookiee is their “shag carpet” appearance: a trait that in conjunction with their exceptional height makes this native Kashyyyk species immediately recognizable. Notably, visibility is an attribute not conducive to the Dark side’s desired covert operations. W. wookiee as a subspecies are identifiable by their fur, which can be uniform in color or a composite of the colors red, brown, and/or chestnut.
At New Order Laboratories, we aim to identify the biological mechanism responsible for W. wookiee hairiness. In conjunction with the Cloning Division, we hope to eventually create hairless, humanoid Wookiee clones for the Dark side.
To begin, we hypothesize that distinct morphological features, in other words, hairiness are a result of Wookiee specific protein-coding gene(s). As Wookiees are a humanoid species, we postulate that when an all-against-all comparison is done with the Wookiee wookiee and Homo Sapien genomes, a portion of the Wookie genome will be distinct. As a consequence, we suggest that elements of these distinct regions will likely be responsible for the hairiness or uniform coat of hair that is so distinct.
MATERIALS AND METHODS
Systematic analysis of the Wookiee genome
A complete BLASTP search of all human, and Wookiee proteins from Ensembl (Hubbard et al. 2007) v 46 with an E-value threshold of 1 x 10-4 was performed according to the protocol set forth in Knowles and McLysaght (2009). Un-ambiguous orthologs were identified and synteny blocks anchored to these regions. A region of 10 genes or less was deemed an acceptable gap between anchors, and thus variability in the gene order was allowed. Anticipated orthologous ORFs in the regions above were defined as BLAT (Kent 2002) or SSearch (Pearson and Lipman 1988) sequence matches.
Ensembl noted several orthologs in more distantly related humanoid species. Further examination of the annotated orthologs was completed to determine if these were old genes that became inactivated in wookiees. Potential orthologs that contained multiple implausible small introns were discarded. For example, the current Ensembl version proposes a Chiss ortholog of BLU31 (gene responsible for their characteristic blue skin), but further investigation concluded that the “gene” lacks possible introns and thus cannot produce any resemblance of a protein in wookiee. Otherwise, the absence of the phenotypic characteristics of plausible orthologs was attributed to the theorized inactivation within the genomes of other humanoid species.
Initial de novo gene candidates were as follows: 100 had a sequence in the expected human region; 65 had conceivable orthologs in the expected human region; 20 candidates exhibited partial nucleotide similarity; 9 Wookiee genes were deemed artifacts; and one candidate had a possible ortholog in Ewok. A final list of candidates consisted of 5 genes: 4 had uninterrupted ORF in Wookiee of at least 50% of the length of the human ORF.
Multi- PipMaker (Schwartz et al. 2000) was used to align the sequence of the wookiee gene to the syntenic location in the human genome. JalView (Clamp et al. 2004) was employed to visualize the sequences and make manual adjustments when applicable.
Wookiee subjects
Wookiee wookiee (New Order standard) samples were obtained from the Wookiee wookiee clonal collection on planet Mustafar. Specimens used in quantitative RT-PCR experiments were kept under controlled conditions for 3-5 days.
Knockout wookiees
Conditional knockout wookiees were generated via gene targeting in wild-type standard W. wookiee by TaconicArtemis. In situ hybridizations were done on [carbonite]-sectioned material with a dioxygenic-based labelling system. Northern blots were done with 10µg total RNA via denaturing agarose gel electrophoresis.
Microarray analysis
Microarray analysis was done on the Wookiee Genome 430 2.0 array, and probe data from the Imperial standard CEL files were normalized via the MAS5 method with the R-based bioconductor software. The normalized probe set data was searched for differentially expressed genes with the significance analysis of microarrays.
RESULTS AND DISCUSSION
Identification of W. wookiee genes with no protein-coding match in protein database or syntenic human genomic region
Principal sites of synteny were established between the human and W. wookiee genome using unambiguous orthologs or shared regions pinpointed by the BLASTP search hits. Synteny blocks established spanned 82% and 74% of the human and Wookiee genomes, respectively, and 18,505 of the 22,568 human protein-coding genes characterized in Ensembl were identified within the established range.
A region of 10 genes was established as an acceptable gap between blocks due to the expected high gene order conservation between the two species. An additional screen for plausible orthologs was conducted in these regions as probability higher for the noted locations in the genome.
An initial 200 candidate Wookiee proteins were identified (no BLASTP hit in the human genome). For 65 proteins, a plausible ortholog existed but upon further examination with BLAT and SSearch only partial nucleotide similarity was identified, leaving 20 candidates. Wookiee genes with conceivable orthologs in other species were also excluded i.e. ewok ortholog.
For the purposes of this screen, only characterized or known W. wookiee genes by Ensembl were considered. Above protocol resulted in one Wookiee protein-coding gene (GLABR) that did not appear to have an ortholog in any other species; genome, however there did appear to be sequence similarity at the expected location of the gene in humans. It is therefore possible that the GLABR gene is of de novo origin. (GLABR was assigned to the Wookiee protein coding for hairlessness as “glaber,” a Latin word meaning hairless or bald).
In order to confirm the de novo origin of the GLABR ortholog, we performed a multiple sequence alignment of the GLABR sequence with the orthologous human sequence (Figure 1). The critical mutation that allows the production of a protein is the deletion of an A nucleotide in the GLABR ortholog, which is present in the human ortholog. This causes a frameshift in Wookiee that results in a much longer ORF capable of producing a 137-amino-acids-long protein; in contrast, the human sequence has a stop codon after only 78 potential codons.
Figure 1. (Click to Enlarge) Sequence changes in the origin of W. wookiee GLABR from noncoding DNA: multiple sequence alignment of the gene sequence of the W. wookiee gene GLABR and similar nucleotide sequences from the syntenic location in humans. The start codon is indicated on the figure.
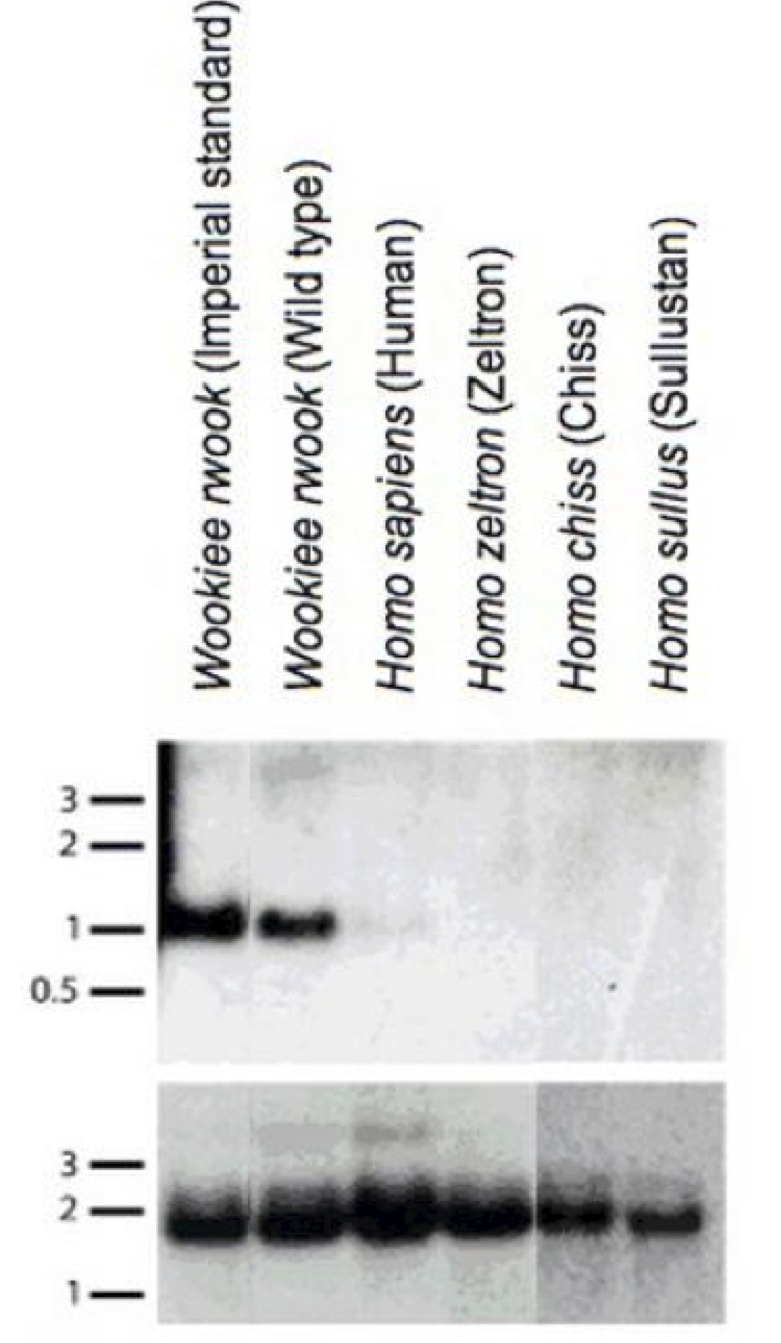
The identified W. wookiee transcript GLABR spans a 453kb region that can be aligned with the human genome, but the human homeolog is free of annotated transcripts or expressed sequence tags (ESTs). We prepared a northern blot with RNA from wookiees and from humans to substantiate the findings that the human aligned region is not expressed (Figure 2); a signal was obtained from the standard W. wookiee subspecies as well as the wild-type W. wookiee species, but not from the human, and other closely related near-human species. This indicates that the non-coding ortholog of the gene is the ancestral sequence, and the gene that confers hairiness to W. wookiees must have originated after the speciation event, approximately 4.7 million years ago. Thus, the gene must have arisen de novo in W. wookiee.
Figure 2. (Click to Enlarge) Northern blot with RNA from Wookiees, human, and related humanoid species. 1kb mRNA fragment containing the putative GLABR sequence is observed only in Wookiee samples, denoting non-expression in other humanoid type gene expression profiles. Note in this figure, the old New Order nomenclature for W. wookiee (W. rwook) is used.
GLABR protein function
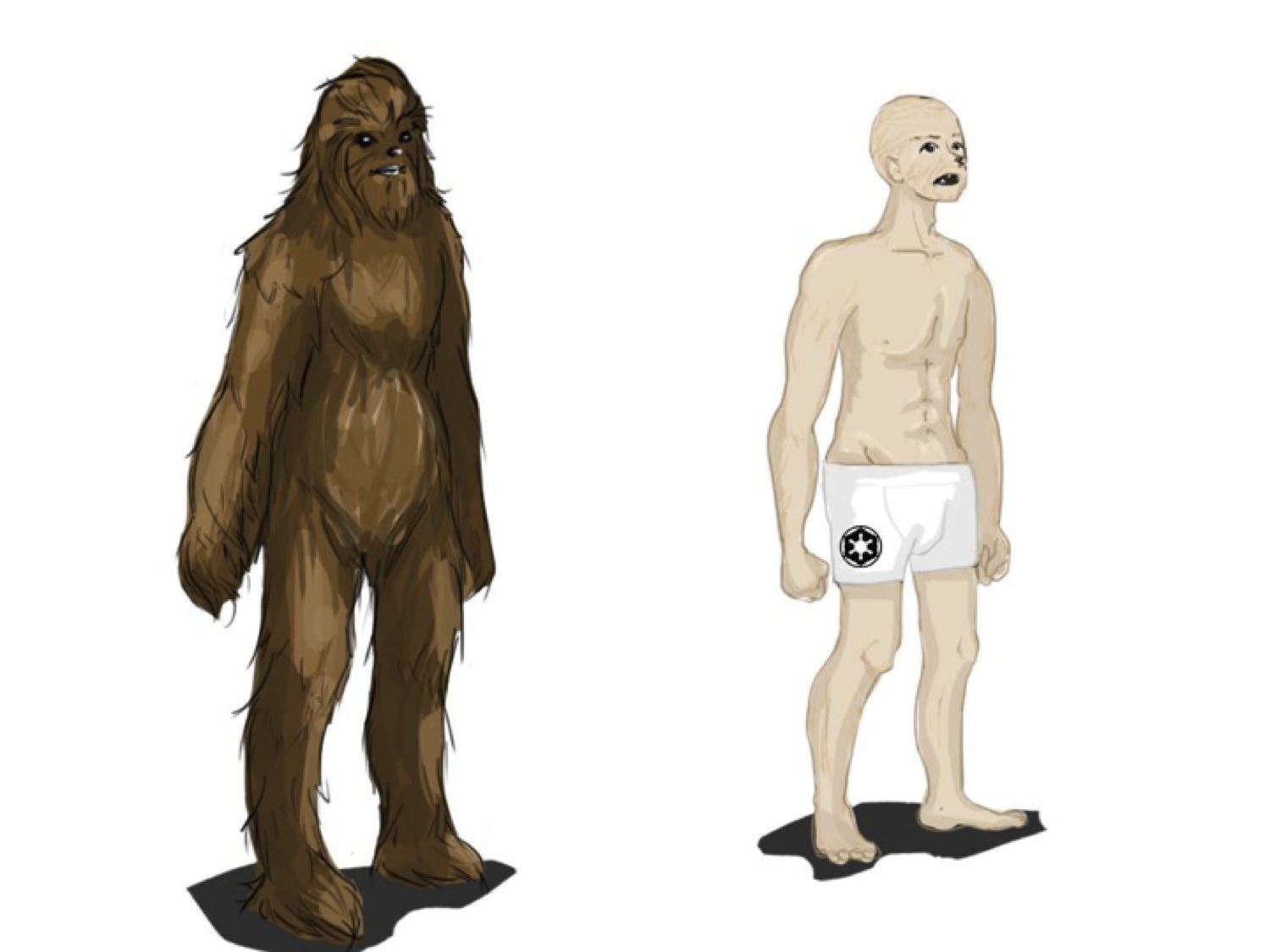
We designed a conditional knockout of the whole gene region to study the function of this GLABR and confirm our hypothesis. We find that the wookiees that lack the GLABR transcript “W. wookiee glaber” are viable and fertile, and the general physiology is not changed, aside from the reduction in androgenic hair growth (Figure 3).
Figure 3: (Click to Enlarge) Artist’s representation of a wild-type wookiee (left) and the GABR variant (right). The latter exhibits a marked reduction in androgenic hair, but the vellous hair is still present.
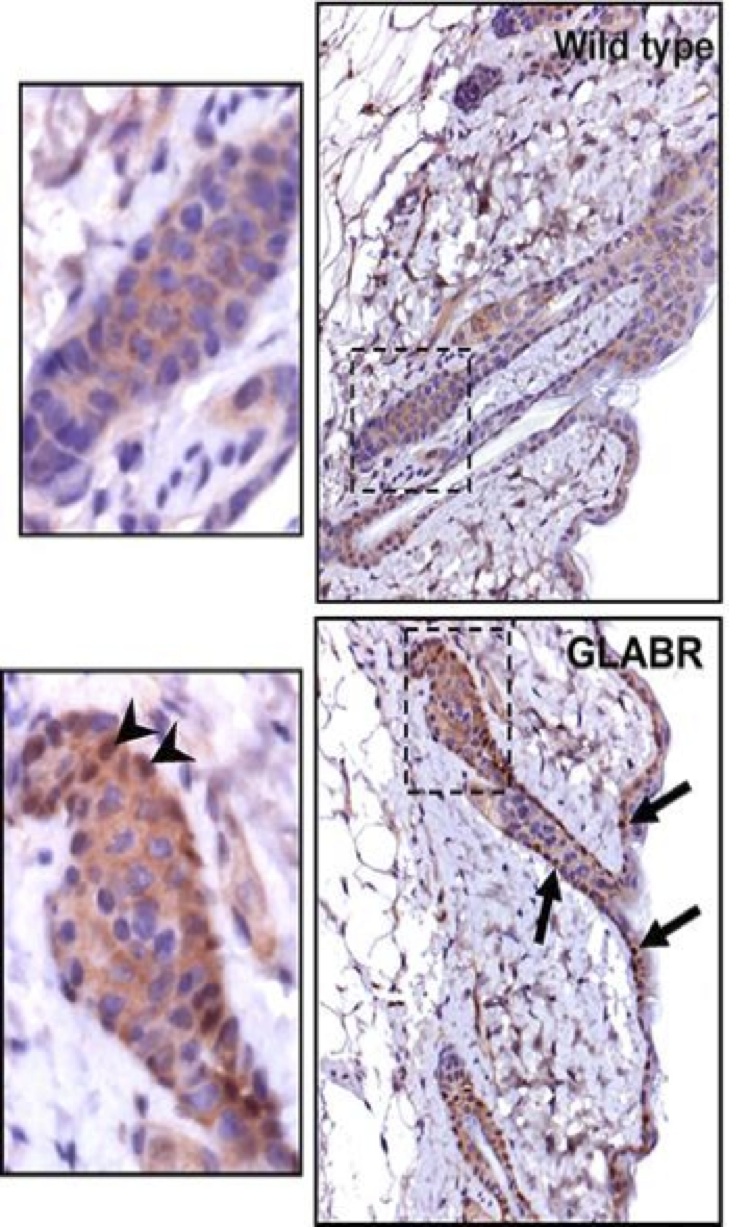
Skin samples were taken from wild-type and GLABR strain wookiees and in situ hybridizations were performed on carbonite-sectioned epithelial material with a dioxygenic-based labeling system (Figure 4). The wild-type section exhibits higher androgen localization towards the androgenic hair roots, whereas the GLABR variant section exhibits much less. These results complement the previous observation that androgenic hair growth was markedly reduced in the GLABR variant (Figure 3). Although for the purpose of this study, it was only necessary to confirm that there was reduced hair growth in the GLABR variants, it would be worth investigating the exact mechanism by which this reduction occurs. Such studies would also examine possible side effects exhibited in the knockout variant.
Figure 4: In situ hybridizations on carbonite-sectioned epithelial specimens from wild-type and GLABR variants, with a dioxygenic-based labeling system.
CONCLUSIONS
This is the first study to focus on uncovering the gene or mechanism behind Wookiee wookiee’s characteristic coat of hair. It is also serves as the first comparative human to W. wookiee genome research. Prior to this research, there are few reports of research into wookiees, and none assessing what genome similarity with humans (if any) could be responsible for their humanoid appearance.
The GLABR protein described in this study functions exclusively to code for W. wookiee’s hairiness. As noted in our results, the hairless wookiee phenotype does not appear to exhibit additional morphological or physiological changes, although we did observe some anecdotal evidence of behavioural modifications. In our knockout W. wookiee glaber (or “Chewie2.0” as we called him), we noted a more docile demeanor when conducting routine laboratory assessments. We hypothesize that the resulting hair loss correlates to a hormonal difference in the normal wookiee androgen levels. This possibility is concerning as it may preclude undesirable physical changes, such as (but not limited to) reductions in muscle mass or physical strength. At present, a research program is being developed to resolve whether this phenomenon is present in all subsequent knockouts.
Further study is also warranted to assess W. wookiee glaber skin sensitivity. While not usually an apparent issue in the species, it is conceivable that rendering them hairless may manifest problematic symptoms. Here, we noted several rash-like symptoms when subjects were clothed, although it remains unclear whether this was a reaction to the garment’s material, or due to other environmental factors such as room temperature or light. W. wookiee culture does not generally encourage individuals to be clothed, but one could argue that such items are deemed culturally unnecessary due to their thick coat of hair. Nevertheless, as seen in mammals, it is likely that W. wookiee hair provides protection from the environment. It is therefore important to weigh the positive and negative impacts of generating a hairless knockout in the context of these various points.
The gene reported in this study is the first case of a W. wookiee protein-coding gene that appears to be restricted to the W. wookiee genome. It is well-documented by this research that the GLABR protein-coding gene is not present in the human genome or any other humanoid species. It is therefore likely that the gene appeared in the W. wookiee genome after the divergence from the human lineage. To conclude, we would hypothesize that Wookiee-specific genes are responsible for Wookiee specific traits i.e. GLABR is responsible for hairiness in Wookiees. It will be interesting going forward to explore whether non-specific genes in Wookiees code for similar characteristics between humanoid species.
Acknowledgments
The authors would like to thank the dark side of the Force, and Conan Motti for donating his life to Science (source of Homo sapien tissue).
Conflict of interest
Dark side.
Funding
The authors gratefully acknowledge the funding support from the New Order and the Dark Lords of the Sith for a studentship awarded to E.M. J.S. was supported through a NOR research scholarship from the Galactic Research Council.
LITERATURE CITED
Clamp M, Cuff J, Searle SM, Barton GJ. (2004) The Jalview Java alignment editor. Bioinformatics 20:426–427.
Heinen, TJ, Staubach F, Haming D, Tautz D (2009) Emergence of a new gene from an intergenic region. Current Biology 19:1527-31.
Hubbard TJ, Aken BL, Beal K, Ballester B, Caccamo M, Chen Y, Clarke L, Coates G, Cunningham F, Cutts T, et al. (2007) Ensembl 2007. Nucleic Acids Res 35:D610–D617.
Kent WJ. (2002) BLAT—the BLAST-like alignment tool. Genome Res 12:656–664.
Knowles, DG, McLysaght A. (2009) Recent de novo origin of human protein-coding genes.
Genome Res 19:1752-9.
Pearson WR, Lipman DJ. (1988) Improved tools for biological sequence comparison. Proc Natl Acad Sci 85:2444–2448.
Schwartz S, Zhang Z, Frazer KA, Smit A, Riemer C, Bouck J, Gibbs R, Hardison R, Miller W. (2000) PipMaker—a web server for aligning two genomic DNA sequences. Genome Res 10:577–586.