THE MACGYVER PROJECT: GENOMIC DNA EXTRACTION AND GEL ELECTROPHORESIS EXPERIMENTS USING EVERYDAY MATERIALS

Abstract:
DNA extraction and separation by agarose gel electrophoresis is a simple and exciting process that anyone can perform. However, the high cost of specialized equipment and chemicals often hinder such an experiment from being carried by members of the high school community. Here, we describe a cost effective way of extracting and electrophoresing DNA under a prescribed MacGyver limitation – that is using only materials available from a grocery store or shopping mall.
In order to carry out this project, we decided to first divide the procedure into three specific sections, each to be addressed individually. Doing this, you find that the following challenges are present. They are: (i) extraction of DNA, (ii) gel electrophoresis of DNA and (iii) visualization of DNA.
Extraction of DNA in a Research Setting:
In a conventional research setting, the first step in extracting DNA involves breaking open the cell’s membrane by using physical or chemical means. Examples include the use of sonication (sonic waves), homogenization equipment (blender, french press, mortar and pestle) or selective detergent use. To us, exploration of detergents (or soaps) or physical steps (mortar and pestle) were the most obvious choices.
Once the cell/tissue has been lysed, usually subsequent steps involve an attempt to purify or enrich the sample for your DNA (i.e. get rid of the other stuff). This can be done in a number of ways, many of which are not technically feasible under our MacGyver rubric. However, a cell lysate can be easily enriched for DNA using alcohol to selectively precipitate DNA, forming it to clump into a “snot” like entity. Consequently, we have decided to focus on the extraction of genomic DNA since this type of sample works best under the alcohol precipation protocol and is also the most abundant DNA species for ease of visualization at both the extraction and gel steps.
A genomic prep is also interesting from an educational point of view. A teacher could pose ‘why do people want to obtain genomic DNA?’ – a question which can lead to numerous discussions pertaining to the cloning of new genes, organisms (Dolly the Sheep); having a source of DNA for fingerprinting purposes (CSI, forensics), or even the preparation of a sample for the sequencing of the organism’s entire genome (Human Genome Project). Focusing on genomic DNA also allows a student to explore samples from different sources, where there is a reasonable possibility that the differences in genome size can be discerned. As a bonus, the precipitation of DNA into a ‘snot’ like form adds an added ‘wow’ factor to the activity.
MacGyver Extraction of DNA:
Reagents and materials for this portion of the experiment are fairly straightforward. Here, one needs to find a tissue source, some manner of physical breaking, clean water/buffer solution, a soap, and an alcohol. In addition, some type of container to do this in will be required, preferably one that is transparent in nature.
Tissue Source:
Although the chemical characteristics of the DNA isolated is practically identical from organism to organism, the cell in which it is housed can vary greatly. Consequently, choice of tissue is the largest variable, but arguably the best element for a student to play around with. We have tried extracting DNA from onion, split peas, corn, yeast, bean sprouts, wheat germ, kiwi, banana and even human cheek cells, all with varying degrees of success. We find that overall, good DNA can be achieved with any fresh produce or grain material, as long as the experimenter is willing to play around to find optimal conditions. Overall, however, we recommend onion or banana tissue as the best sources of low maintenance DNA extraction. We also recommend wheat germ, in that a “fast” procedure exists that works very well. Finally, DNA isolated from a student’s own cheek cells is an interesting alternative, since it brings in a personal element to the activity. However, it should be noted that isolation from cheek cells is less reliable.
Physical Step:
Depending on the tissue you use, a physical step may be warranted. This could be as involved as cutting up the tissue and then using a mortar and pestle, or as minor as banging around with a wooden chopstick. In general, plant material and meat cuts would benefit from such a step, since plants have a tough cell wall to contend with, and meat usually contains tough muscle striations.
Buffer Solution:
This is the fluid that the sample needs to be immersed in, and as such has the role of simply keeping your DNA safe. There are many possible MacGyver type buffers that one can use for the extraction procedure, which all basically work (see Table 1). We should note, however, that distilled or bottled water should be used. Tap water appears to have a minor effect on the extraction procedure, and is actually very detrimental in the subsequent gel steps described later.
TABLE 1: BUFFER RECIPES*
A: “Saline Buffer”
0.9% saline (contact lens solution)
B: “Regular Buffer”
1.5g table salt (NaCl)
5.0g baking soda (sodium bicarbonate)
to a final volume of 120ml with water
C: “Acidic Buffer”
1.5g table salt (NaCl)
5.0g baking soda (sodium bicarbonate)
5mL of vinegar
to a final volume of 120ml with water
D: “Proteinase Buffer”
1.5g table salt
5.0g baking soda
2.5mL Complete eye solution or 2 protein tablets (Complete brand)
to a final volume of 120ml with water
E: “Meat Tenderizer Buffer”
100mL hot water
3g meat tenderizer
*Note that water alone can also be used effectively.
Soap/Detergent:
Liquid dish washing soap was generally used. In terms of specifics, we found better results when the soap was colourless, and found that unscented versions may work best overall. Essentially, most soaps available at grocery stores have additives that play roles in preservation, scent, colour, stain removal etc, etc. Although these things are usually not an issue with the technique, choosing the simplest soap is a good thing to troubleshoot with if you are not having success with your extraction. In general, we found that using about 5ml (or less) of detergent per 120ml solution works well.
Filtering:
Depending on your tissue choice and also the amount of tissue you have decided to use, it may be pertinent to include a filtering step to remove the debris. This allows a better visualization of the DNA snot effect at the end. Filtering can be easily accomplished using coffee filters, or cheese cloth. Note that when using cheese cloth, you will need use 3-4 layers of cheesecloth. Coffee filters are generally easier because they are cheaper, more accessible, and easier to cleanup.
Precipitation with Alcohol:
Precipitation of DNA is done by contact with alcohol. This alcohol is available at pharmacies as rubbing alcohol of the 99% iso-propanol variety. Alternatively, most schools will have access to 95% ethanol stocks. The alcohol should be added in a layering manner so that one can see the DNA forming at the interface between your sample and the alcohol. You should add at least two times the volume of your original sample. If you do not see “snot” forming, then mixing everything together should get the desired effect. Note that if you intend to resort to mixing, then the filtering step may be especially necessary for ease of visualization.
DNA precipitates under alcohol treatment because it is naturally hydrophobic and as such will tend to clump together if the solvent is not optimal (i.e. water). As a result, adding an alcohol to your prep will mean that DNA is not completely solvated in its optimal environment (water), and will therefore aggregate, and precipitate out of solution. It should be stressed that the degree and ease of clumping will depend on the amount of DNA present, and also the concentration at which it exists. Therefore, if you see minimal amounts of DNA, you should be able to correct this by (i) trying to isolate from more tissue, and (ii) resuspend the material in a smaller volume of fluid.
TABLE 2: SUGGESTED MACGYVER GENOMIC DNA EXTRACTION PROTOCOLS:
GENERAL PROTOCOL
1. If necessary, slice up DNA source of choice. Use an amount about the size of a strawberry.
2. Using a mortar and pestle, grind up sample while gradually adding 10mL of prepared buffer solution with the detergent already added. Grind for at least 5 minutes with all of your weight and strength to ensure that you break open the cell membrane and reach a creamy soup consistency. If the sample is too thick after grinding, add more saline solution to achieve the optimal thickness so that the liquid portion of the sample is able to pass through the filter, while the larger cellular material remains behind on the filter. Note: If the DNA sample is frozen, it is considerably easier to grind.
3. Filter your sample’s juice into a small beaker. Let the solution drip into the beaker until all of the liquid has passed through the filter. If this takes too long, simply squeeze all of the juice from the sample through the filter.
4. Add 2 volumes (this means approximately two times the volume of the sample present) of ice cold alcohol down the side of the beaker with a straw or pasteur pipette. Do this step slowly to enable the alcohol to form a layer on top of the juice layer. As you let the beaker sit, the DNA should precipitate. The longer you wait, the more DNA you should see. If you don’t see precipitation, gently mix everything together.
5. The DNA precipitates should resemble a thread of translucent white snot,at the interface between the juice and alcohol. After a considerable amount of time, the DNA may eventually float to the top of the alcohol layer.
6. You can remove the DNA with a wooden popsicle stick or glass rod. DNA adheres well to the wood.
QUICK PROTOCOL
1. Place a teaspoon or less of wheat germ in your cup.
2. Add about 10ml distilled water and crush gently with a popsicle stick/chopstick for 1 minute.
3. Then add a squirt of dish detergent and crush gently with popsicle stick for 2 minutes.
4. Slowly pour alcohol down the length of the stir stick, layering the alcohol on top of the water.
5. Set the container down on a table and look for the DNA at the interface between the alcohol and water.
CHEEK CELL PROTOCOL
1. Measure 10 ml of “regular buffer (from Table 1), pour buffer in mouth and swirl around cheeks for about 1 minute.
2. Spit the water back into a container, preferably something relatively narrow like a test tube.
3. Squirt a bit of liquid soap to the sample and mix well with popsicle stick. If you can mix by inversion, then do so gently about 20 times.
4. Add 10 ml cold alcohol slowly to the sample and make sure to pour it at an angle down the side of the test tube so that two layers are formed. Do this very gently, with a straw, etc. It’s important that the two layers are not disturbed.
5. Wait for about 10 minutes and the DNA will appear afloat on the alcohol layer.
Gel Electrophoresis of DNA in a Research Setting:
Electrophoresis is a way of separating molecules based on charge and size. In our case, we want to separate different sizes of genomic DNA molecules obtained from fruits, vegetables and yeast. Generally, polysaccharide polymers such as agarose or acrylamide are used to form the electrophoresis gels. Because DNA is negatively charged, one can force it to travel through the gel by applying an electric field in the system. Normally, this is achieved by using special gel apparatus designed to facilitate the production or casting of the “gel” as well as allow a platform to immerse the gel in an ion containing buffer to create an electric field. Such an apparatus will run a minimum of about $500, and power packs normally used to deliver ~80V of voltage can run in a similar price range. Consequently, this aspect of the MacGyver project is arguably focused on cost savings and concentrates primarily on finding a substitute for agarose and directions for producing a gel apparatus.
MacGyver Gel Electrophoresis of DNA:
Gel Material:
Agarose is a component of seaweed and as such is a refined purified molecule derived from a common food thickener known as “Agar Agar.” which incidentally can be easily found at most oriental style food stores. “Agar Agar” can be purchased in either a flake, noodle or powder form. Try to ensure that it does not contain any additional ingredients (such as glucose) as these ingredients may interfere with the formation of the gel matrix. To make the gel, it is recommended to use “Agar Agar” in the powder form rather than the flake or noodle form. The other larger forms tend to require cutting and additional filtering which is problematic and very messy.
Running Buffer:
You will need to prepare a running buffer which is required to make the gel, and also required as the fluid that will ultimately immerse your solidified gel to allow the electric field to be conducted. The Macgyver running buffer recipe is as follows:
– 0.05g of NaCl (this is the principle ion)
– 2g of Baking Soda (Sodium Bicarbonate)
– bring to 1L with distilled bottle water
pH’d using pet store aquarium pH kit to approximately pH7.5. (we used alkaline buffer made by Seqchem).
Gel Apparatus:
Although a research grade gel box is costly, it is in fact a relatively simple piece of equipment. In essence it is a large container (let’s call it a buffer chamber) that can hold fluid, whereby opposite ends are connected to a power source setting up a positive/negative electrode scenario. The buffer chamber needs to be able to conveniently hold a smaller container (let’s call this the gel casting chamber) that is used to make and hold the gel. Furthermore, the smaller gel casting chamber needs to fit in such a manner as to be in the middle between the two electrodes of the buffer chamber.
Assembling this electrophoresis box should be quite straightforward and even enjoyable for those that like to “make things.” We have included here a cartoon guide (Figure 1) for making these two chambers out of common plasticware (tupperware, soap dish, etc). For the electrode connections, we were limited to stainless steel screws (5cm) and stainless steel wire (20 gauge), which worked fine. However, a more elaborate electrode system would be easy to make with a visit to a proper hardware store.
As an alternative to using tupperware, we have also found Lego to be extremely useful in custom making a buffer chamber to fit your gel casting chamber. Note that Lego chambers will need to be lined with a water tight material, but recently Glad has developed a new “Glad Press’N Seal” material which works perfectly and is easy to handle.
Finally, a “comb: will need to be made. This is a contraption that allows small wells to be formed in your gel. Here, we found lego to be especially useful, but in a bind, we also made combs by cutting out pieces of plastic, or taping the teeth of a real hair comb.
FIGURE 1: MACGYVER GEL BOX CONSTRUCTION NOTES.
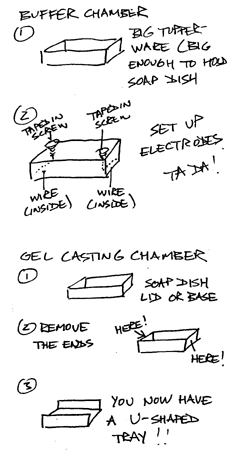
Gel Preparation:
Using the powdered “Agar Agar” you will want to make a 1.2% to 1.5% gel (w/v or 1.2g to 1.5g per 100ml of running buffer). In a separate container, (a flask for instance) weigh out the required amount of “Agar Agar” and add the required amount of running buffer (specific amounts will be dictated by the size of your gel casting set-up). The flask should be large enough so that the “Agar Agar” solution forms a 2 cm layer at the base. This is required to ensure a large surface area in contact with the hotplate for effective melting. It is recommended to use a hot plate and continued stirring to heat the agar solution evenly. An alternative is to use a microwave, but with this method one must use a low heat level at short intervals while swirling the mixture frequently. If this procedure is not carried out carefully, the solution will likely overboil and create a large mess. Note that if the “Agar Agar” solution is not completely dissolved, this will greatly affect the mobility of the DNA as it travels through the gel. Furthermore, undissolved bits of agar will be stained during the staining procedure, making it difficult to view the bands of DNA.
Note: some schools have access to the “Agar” used to make bacterial plates. This material works very well in pouring gels (much better than “Agar Agar”) We recommend making a 1% w/v gel for genomic visualization purposes.
Make sure that you have sealed the open ends of your casting gel chamber with masking tape or similar material. Pour the melted mixture into your casting gel chamber. Remember to also insert the “comb” so that well formation can occur. It will take approximately 20 to 30 minutes at room temperature for the gel to solidify. During this time do not disturb the gel. The gel is quite delicate, so be very careful when you remove the comb before loading your sample
The gel should be poured to about a 0.5cm to 1.0cm thickness. Thicker gels allow the opportunity to make deeper wells, thereby allowing a larger sample to be loaded. Thinner gels should run faster. Note that if the gel is too thin, it may float when immersed. Try to use the gel as soon as possible. Although you can wrap it in plastic wrap and store it in the fridge overnight, they will work better when used fresh.
DNA Sample Preparation:
Your genomic snot DNA can be removed at the end of the extraction procedure using a wooden stir stick. The DNA should then be dipped in a 70% alcohol solution which facilitates removal of excess salts. This is very important for effective dissolving as well as effective gel running. The DNA can then be air dried for approximately 10 minutes to evaporate residual alcohol, and then dislodged into a small volume of running buffer (the smaller the better, i.e. <0.5mls). One should note that genomic DNA can often take days to dissolve properly. We suggest allowing the DNA to dissolve overnight at room temperature (if you have access to temperature up to 50C, then that is best). Be gentle to your sample as the large genomic DNA is very prone to shearing. Once the overnight dissolving step is finished, consider this your DNA sample regardless of whether it has dissolved to completion.
Your sample will then need to be treated with a loading buffer. This is essentially something that will cause your sample to be viscous so that it can indeed “sink” into your wells during loading. Often, a loading buffer also has a dye, which will travel in the same direction as the DNA, and also makes the gel loading easier to see.
Our recipe is as follows:
– 0.5ml glycerol/glycerine (available at the pharmacy section)
– 0.1ml distilled water
– several drops of Club House Red Food Colouring (note that choice of dye can affect outcome greatly – we didn’t have a lot of success finding something good here, so a last resort would be to use without a dye).
This loading buffer can be used as 5X to 10X meaning that for 0.5ml of sample, you would need to add 0.05 to 0.1ml of loading buffer. Mix carefully. Your sample is now ready for the gel.
Gel Running.
Place your solidified gel (in gel casting chamber) within the larger buffer chamber. Add running buffer until the gel is immersed such that there is at least 3mm of fluid above the gel. Keep in mind that the more fluid you add, the slower the gel will run.
To your wells, load as much sample (with loading buffer) as possible. This is probably best done with a plastic stir stick attached to some type of rubber bulb. Essentially you would like to assemble something that can deliver fluid through a small tube. Once all the samples have been loaded, you will want to connect the electrodes to a series of 9V batteries. Your DNA is negatively charged so you want to position the positive electrode at the end “away” from the wells. Basically, one battery will suffice but will be very very slow (overnight run scenario). 5 to 7 or them lined up in a series circuit should deliver a good amount of voltage. Note that when the gel is running, DO NOT stick your finger in the fluid. Depending on the number of batteries you use, as much as 100mAmps of current is delivered (enough to give you a small shock).
When the circuit is running, a good visual check is to see that bubbles are forming from the wire electrodes, and usually most visible at the positive end.
The optimal amount of time to run the gel is, frankly, something that is difficult to predict, as it depends on the size of your gel, the thickness of your gel, the amount of running buffer in the system, the amount of voltage applied, and even the wiring set-up used. Consequently, this is the one thing where you will definitely have to play around. However, with a 5 to 7 battery set-up, you will need at least a minimum of 1hour, and possibly more. In addition, if differentiating genomic preps is in order (i.e. different genome size), you may need to experiment further with run time to see this potential difference.
Visualizing DNA in a Research Setting:
Upon completion of electrophoresis, the location of the bands need to be visualized. A common way to detect the bands is to stain the gel with ethidium bromide, which fluoresces under ultraviolet light to view the DNA. Unfortunately, both ethidium bromide and ultraviolet light are hazardous (Ethidium is higly carcinogenic and UV light can burn a person’s eyes without proper safety measures) and are therefore, not ideal for use among high school students.
MacGyver Way to Visualize the DNA:
To visualize the DNA bands, the gel was stained in a Methylene Blue solution. Methylene Blue consists of the salt methylene blue chloride. In water, the salt disassociates into a positively charged methylene blue ion that is colored blue and a negatively charged chloride ion, which is colorless. This blue chromophore is then able to bind to the positively charged DNA in the gel. Methylene blue is a convenient stain to use in the lab because it is chemically safe, reusable, and detects the presence of more than 20ng DNA/band.
More importantly, methylene blue can conveniently be found at most pet stores. It is generally sold as an aquarium disinfectant at a 5% concentration. The best gel staining results were obtained immersing the finished gel in a 0.02% methylene blue (in distilled water) solution overnight at room temperature. Using this protocol, destaining excess the excess colour with distilled water incubations was not required. We found that higher concentration solutions tended to stain the gel too dark making it difficult to differentiate the background from the DNA bands. This was observed even after destaining with distilled water. Also, if the “Agar Agar” powder was not completely dissolved in the gel, the non-dissolved flakes were stained dark blue, preventing the formation of distinct, visible bands.
FIGURE 2: CORN GENOMIC DNA PREP MACGYVER STYLE.

Conclusions:
Overall, we have described an effective outline of methods to perform some basic molecular biology techniques under the MacGyver limitation (Figure 2). We should stress however that doing this in your own classroom will inevitably require some working out of your own. This is due to a multitude of considerations such as “Do I have enough DNA?” “what is my gel set-up” “what type of specialized reagents do I have at my disposal to make things even more efficient?” etc. However, it is hoped that the information presented here will make your troubleshooting as smooth as possible. Good luck!